Supported by Dr. Osamu Ogasawara and  providing providing  . . |
|
Last data update: 2014.03.03 |
Browse Objects in EnvironmentDescriptionThe Usage
browseEnv(envir = .GlobalEnv, pattern,
excludepatt = "^last\.warning",
html = .Platform$GUI != "AQUA",
expanded = TRUE, properties = NULL,
main = NULL, debugMe = FALSE)
Arguments
DetailsVery experimental code: displays a static HTML page on all platforms
except Only allows one level of recursion into object structures. It can be generalized. See sources for details. Most probably, this should rather work through using the tkWidget package (from https://www.bioconductor.org). See Also
Examples
if(interactive()) {
## create some interesting objects :
ofa <- ordered(4:1)
ex1 <- expression(1+ 0:9)
ex3 <- expression(u, v, 1+ 0:9)
example(factor, echo = FALSE)
example(table, echo = FALSE)
example(ftable, echo = FALSE)
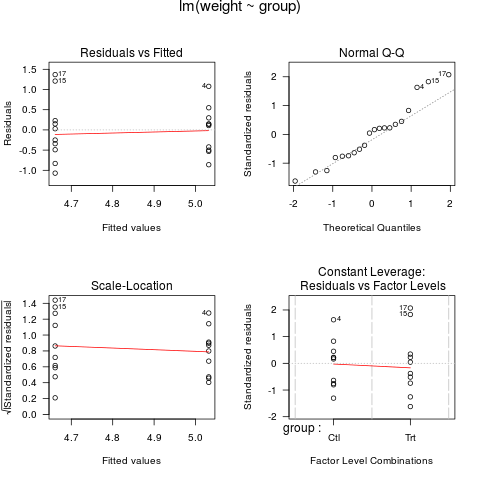
example(lm, echo = FALSE, ask = FALSE)
example(str, echo = FALSE)
## and browse them:
browseEnv()
## a (simple) function's environment:
af12 <- approxfun(1:2, 1:2, method = "const")
browseEnv(envir = environment(af12))
}
Results
R version 3.3.1 (2016-06-21) -- "Bug in Your Hair"
Copyright (C) 2016 The R Foundation for Statistical Computing
Platform: x86_64-pc-linux-gnu (64-bit)
R is free software and comes with ABSOLUTELY NO WARRANTY.
You are welcome to redistribute it under certain conditions.
Type 'license()' or 'licence()' for distribution details.
R is a collaborative project with many contributors.
Type 'contributors()' for more information and
'citation()' on how to cite R or R packages in publications.
Type 'demo()' for some demos, 'help()' for on-line help, or
'help.start()' for an HTML browser interface to help.
Type 'q()' to quit R.
> library(utils)
> png(filename="/home/ddbj/snapshot/RGM3/R_rel/result/utils/browseEnv.Rd_%03d_medium.png", width=480, height=480)
> ### Name: browseEnv
> ### Title: Browse Objects in Environment
> ### Aliases: browseEnv wsbrowser
> ### Keywords: interface
>
> ### ** Examples
>
> #if(interactive()) {
> ## create some interesting objects :
> ofa <- ordered(4:1)
> ex1 <- expression(1+ 0:9)
> ex3 <- expression(u, v, 1+ 0:9)
> example(factor, echo = FALSE)
> example(table, echo = FALSE)
d
b A B C D E
A 10 . . . .
B . 10 . . .
C . . 10 . .
> example(ftable, echo = FALSE)
> example(lm, echo = FALSE, ask = FALSE)
> example(str, echo = FALSE)
int [1:12] 1 2 3 4 5 6 7 8 9 10 ...
function (name, pos = -1L, envir = as.environment(pos), all.names = FALSE,
pattern, sorted = TRUE)
function (name)
'data.frame': 39 obs. of 5 variables:
$ y : Time-Series from 1962 to 1972: 8.79 8.79 8.81 8.81 8.91 ...
$ lag.quarterly.revenue: num 8.8 8.79 8.79 8.81 8.81 ...
$ price.index : num 4.71 4.7 4.69 4.69 4.64 ...
$ income.level : num 5.82 5.83 5.83 5.84 5.85 ...
$ market.potential : num 13 13 13 13 13 ...
function (object, ...)
List of 18
$ double.eps : num 2.2204460492503130808e-16
$ double.neg.eps : num 1.1102230246251565404e-16
$ double.xmin : num 2.2250738585072013831e-308
$ double.xmax : num 1.7976931348623157081e+308
$ double.base : int 2
$ double.digits : int 53
$ double.rounding : int 5
$ double.guard : int 0
$ double.ulp.digits : int -52
$ double.neg.ulp.digits: int -53
$ double.exponent : int 11
$ double.min.exp : int -1022
$ double.max.exp : int 1024
$ integer.max : int 2147483647
$ sizeof.long : int 8
$ sizeof.longlong : int 8
$ sizeof.longdouble : int 16
$ sizeof.pointer : int 8
List of 4
$ coefficients: Named num [1:2] 1.18e-15 1.00
..- attr(*, "names")= chr [1:2] "Intercept" "X"
$ residuals : num [1:9] -9.01e-16 1.72e-15 -2.47e-16 -2.25e-16 -2.03e-16 ...
$ intercept : logi TRUE
$ qr :List of 6
..$ qt : num [1:9] -1.50e+01 7.75 -2.22e-16 0.00 2.22e-16 ...
..$ qr : num [1:9, 1:2] -3 0.333 0.333 0.333 0.333 ...
.. ..- attr(*, "dimnames")=List of 2
.. .. ..$ : NULL
.. .. ..$ : chr [1:2] "Intercept" "X"
..$ qraux: num [1:2] 1.33 1.26
..$ rank : int 2
..$ pivot: int [1:2] 1 2
..$ tol : num 1e-07
..- attr(*, "class")= chr "qr"
List of 4
$ coefficients: Named num [1:2] 1.18e-15 1.00
..- attr(*, "names")= chr [1:2] "Intercept" "X"
$ residuals : num [1:9] -9.01e-16 1.72e-15 -2.47e-16 -2.25e-16 -2.03e-16 ...
$ intercept : logi TRUE
$ qr :List of 6
..- attr(*, "class")= chr "qr"
List of 4
$ coefficients: Named num [1:2] 1.18e-15 1.00
..- attr(*, "names")= chr [1:2] "Intercept" "X"
$ residuals : num [1:9] -9.01e-16 1.72e-15 -2.47e-16 -2..
$ intercept : logi TRUE
$ qr :List of 6
..$ qt : num [1:9] -1.50e+01 7.75 -2.22e-16 0.00 2.22e..
..$ qr : num [1:9, 1:2] -3 0.333 0.333 0.333 0.333 ...
.. ..- attr(*, "dimnames")=List of 2
.. .. ..$ : NULL
.. .. ..$ : chr [1:2] "Intercept" "X"
..$ qraux: num [1:2] 1.33 1.26
..$ rank : int 2
..$ pivot: int [1:2] 1 2
..$ tol : num 1e-07
..- attr(*, "class")= chr "qr"
List of 4
$ coefficients: Named num [1:2] 1.18e-15 1.00
..- attr(*, "names")= chr [1:2] "Intercept" "X"
$ residuals : num [1:9] -9.01e-16 1.72e-15 -2.47e-16
-2.25e-16 -2.03e-16 ...
$ intercept : logi TRUE
$ qr :List of 6
..$ qt : num [1:9] -1.50e+01 7.75 -2.22e-16 0.00 2.22e-16
...
..$ qr : num [1:9, 1:2] -3 0.333 0.333 0.333 0.333 ...
.. ..- attr(*, "dimnames")=List of 2
.. .. ..$ : NULL
.. .. ..$ : chr [1:2] "Intercept" "X"
..$ qraux: num [1:2] 1.33 1.26
..$ rank : int 2
..$ pivot: int [1:2] 1 2
..$ tol : num 1e-07
..- attr(*, "class")= chr "qr"
List of 60
$ CBoundsCheck : logi FALSE
$ HTTPUserAgent : chr "R (3.3.1 x86_64-pc-linux-gnu x86_64 linux-gnu)"
$ OutDec : chr "."
$ add.smooth : logi TRUE
$ bitmapType : chr "cairo"
$ browser : chr "/usr/bin/firefox"
$ browserNLdisabled : logi FALSE
$ check.bounds : logi FALSE
$ citation.bibtex.max : int 1
$ continue : chr "+ "
$ contrasts : Named chr [1:2] "contr.treatment" "contr.poly"
..- attr(*, "names")= chr [1:2] "unordered" "ordered"
$ defaultPackages : chr [1:6] "datasets" "utils" "grDevices" "graphics" ...
$ demo.ask : chr "default"
$ deparse.cutoff : int 60
$ device :function (file = if (onefile) "Rplots.pdf" else "Rplot%03d.pdf", width,
height, onefile, family, title, fonts, version, paper, encoding, bg,
fg, pointsize, pagecentre, colormodel, useDingbats, useKerning, fillOddEven,
compress)
$ device.ask.default : logi FALSE
$ digits : int 7
$ dvipscmd : chr "dvips"
$ echo : logi TRUE
$ editor : chr "vi"
$ encoding : chr "native.enc"
$ example.ask : chr "default"
$ expressions : int 5000
$ help.search.types : chr [1:3] "vignette" "demo" "help"
$ help.try.all.packages: logi FALSE
$ internet.info : int 2
$ keep.source : logi FALSE
$ keep.source.pkgs : logi FALSE
$ locatorBell : logi TRUE
$ mailer : chr "mailto"
$ max.print : int 99999
$ menu.graphics : logi TRUE
$ na.action : chr "na.omit"
$ nwarnings : int 50
$ pager : chr "/home/ddbj/local/lib64/R/bin/pager"
$ papersize : chr "a4"
$ pdfviewer : chr "/usr/bin/xdg-open"
$ pkgType : chr "source"
$ printcmd : chr "lpr"
$ prompt : chr "> "
$ repos : Named chr "@CRAN@"
..- attr(*, "names")= chr "CRAN"
$ rl_word_breaks : chr " \t\n"\'`><=%;,|&{()}"
$ scipen : num 0
$ show.coef.Pvalues : logi TRUE
$ show.error.messages : logi TRUE
$ show.signif.stars : logi TRUE
$ showErrorCalls : logi TRUE
$ str :List of 3
..$ strict.width: chr "no"
..$ digits.d : int 3
..$ vec.len : int 4
$ str.dendrogram.last : chr "`"
$ stringsAsFactors : logi TRUE
$ texi2dvi : chr "/usr/bin/texi2dvi"
$ timeout : num 60
$ ts.S.compat : logi FALSE
$ ts.eps : num 1e-05
$ unzip : chr "/usr/bin/unzip"
$ useFancyQuotes : logi TRUE
$ verbose : logi FALSE
$ warn : num 0
$ warning.length : int 1000
$ width : int 80
List of 72
$ xlog : logi FALSE
$ ylog : logi FALSE
$ adj : num 0.5
$ ann : logi TRUE
$ ask : logi FALSE
$ bg : chr "white"
$ bty : chr "o"
$ cex : num 1
$ cex.axis : num 1
$ cex.lab : num 1
$ cex.main : num 1.2
$ cex.sub : num 1
$ cin : num [1:2] 0.15 0.2
$ col : chr "black"
$ col.axis : chr "black"
$ col.lab : chr "black"
$ col.main : chr "black"
$ col.sub : chr "black"
$ cra : num [1:2] 10.8 14.4
$ crt : num 0
$ csi : num 0.2
$ cxy : num [1:2] 0.0597 0.1917
$ din : num [1:2] 6.67 6.67
$ err : int 0
$ family : chr ""
$ fg : chr "black"
$ fig : num [1:4] 0 1 0 1
$ fin : num [1:2] 6.67 6.67
$ font : int 1
$ font.axis: int 1
$ font.lab : int 1
$ font.main: int 2
$ font.sub : int 1
$ lab : int [1:3] 5 5 7
$ las : int 0
$ lend : chr "round"
$ lheight : num 1
$ ljoin : chr "round"
$ lmitre : num 10
$ lty : chr "solid"
$ lwd : num 1
$ mai : num [1:4] 1.02 0.82 0.82 0.42
$ mar : num [1:4] 5.1 4.1 4.1 2.1
$ mex : num 1
$ mfcol : int [1:2] 1 1
$ mfg : int [1:4] 1 1 1 1
$ mfrow : int [1:2] 1 1
$ mgp : num [1:3] 3 1 0
$ mkh : num 0.001
$ new : logi FALSE
$ oma : num [1:4] 0 0 0 0
$ omd : num [1:4] 0 1 0 1
$ omi : num [1:4] 0 0 0 0
$ page : logi TRUE
$ pch : int 1
$ pin : num [1:2] 5.43 4.83
$ plt : num [1:4] 0.123 0.937 0.153 0.877
$ ps : int 12
$ pty : chr "m"
$ smo : num 1
$ srt : num 0
$ tck : num NA
$ tcl : num -0.5
$ usr : num [1:4] -0.58 1.58 -2.09 2.54
$ xaxp : num [1:3] -0.5 1.5 4
$ xaxs : chr "r"
$ xaxt : chr "s"
$ xpd : logi FALSE
$ yaxp : num [1:3] -2 2 4
$ yaxs : chr "r"
$ yaxt : chr "s"
$ ylbias : num 0.2
chr [1:12] "a" "b" NA NA NA "f" "g" "h" "i" "j" "k" ...
List of 2
$ a: chr "A"
$ L:List of 100
..$ : int 1
..$ : int 2
..$ : int 3
..$ : int 4
..$ : int 5
..$ : int 6
..$ : int 7
..$ : int 8
..$ : int 9
.. [list output truncated]
chr "abcdefghijklmnopqrstuvwxyzabcdefghijklmnopqrstuvwxyzabcdefghijklmnopqrstuvwxyzabcdefghijklmnopqrstuvwxyzabcdefghijklmnopqrstuvw"| __truncated__
chr "abcdefghijklmnopqrstuvwxyzabcdefghijklmnopqrstuvwxy"| __truncated__
chr
"abcdefghijklmnopqrstuvwxyzabcdefghijklmnopqrstuvwxyzabcdefghijklmnopqrstu"..
__truncated__
List of 4
$ coefficients: Named num [1:2] 1.18e-15 1.00
..- attr(*, "names")= chr [1:2] "Intercept" "X"
$ residuals : num [1:9] -9.01e-16 1.72e-15 -2.47e-16
-2.25e-16 -2.03e-16 ...
$ intercept : logi TRUE
$ qr :List of 6
..$ qt : num [1:9] -1.50e+01 7.75 -2.22e-16 0.00 2.22e-16
...
..$ qr : num [1:9, 1:2] -3 0.333 0.333 0.333 0.333 ...
.. ..- attr(*, "dimnames")=List of 2
.. .. ..$ : NULL
.. .. ..$ : chr [1:2] "Intercept" "X"
..$ qraux: num [1:2] 1.33 1.26
..$ rank : int 2
..$ pivot: int [1:2] 1 2
..$ tol : num 1e-07
..- attr(*, "class")= chr "qr"
List of 60
$ CBoundsCheck : logi FALSE
$ HTTPUserAgent : chr "R (3.3.1 x86_64-pc-linux-gnu x86_64
linux-gnu)"
$ OutDec : chr "."
$ add.smooth : logi TRUE
$ bitmapType : chr "cairo"
$ browser : chr "/usr/bin/firefox"
$ browserNLdisabled : logi FALSE
$ check.bounds : logi FALSE
$ citation.bibtex.max : int 1
$ continue : chr "+ "
$ contrasts : Named chr [1:2] "contr.treatment"
"contr.poly"
..- attr(*, "names")= chr [1:2] "unordered" "ordered"
$ defaultPackages : chr [1:6] "datasets" "utils"
"grDevices" "graphics" ...
$ demo.ask : chr "default"
$ deparse.cutoff : int 60
$ device :function (file = if (onefile) "Rplots.pdf" else
"Rplot%03d.pdf",
width, height, onefile, family, title, fonts, version,
paper, encoding, bg, fg, pointsize, pagecentre,
colormodel, useDingbats, useKerning, fillOddEven,
compress)
$ device.ask.default : logi FALSE
$ digits : int 7
$ dvipscmd : chr "dvips"
$ echo : logi TRUE
$ editor : chr "vi"
$ encoding : chr "native.enc"
$ example.ask : chr "default"
$ expressions : int 5000
$ help.search.types : chr [1:3] "vignette" "demo" "help"
$ help.try.all.packages: logi FALSE
$ internet.info : int 2
$ keep.source : logi FALSE
$ keep.source.pkgs : logi FALSE
$ locatorBell : logi TRUE
$ mailer : chr "mailto"
$ max.print : int 99999
$ menu.graphics : logi TRUE
$ na.action : chr "na.omit"
$ nwarnings : int 50
$ pager : chr "/home/ddbj/local/lib64/R/bin/pager"
$ papersize : chr "a4"
$ pdfviewer : chr "/usr/bin/xdg-open"
$ pkgType : chr "source"
$ printcmd : chr "lpr"
$ prompt : chr "> "
$ repos : Named chr "@CRAN@"
..- attr(*, "names")= chr "CRAN"
$ rl_word_breaks : chr " \t\n"\'`><=%;,|&{()}"
$ scipen : num 0
$ show.coef.Pvalues : logi TRUE
$ show.error.messages : logi TRUE
$ show.signif.stars : logi TRUE
$ showErrorCalls : logi TRUE
$ str :List of 4
..$ strict.width: chr "wrap"
..$ digits.d : num 3
..$ vec.len : num 4
..$ formatNum :function (x, ...)
$ str.dendrogram.last : chr "`"
$ stringsAsFactors : logi TRUE
$ texi2dvi : chr "/usr/bin/texi2dvi"
$ timeout : num 60
$ ts.S.compat : logi FALSE
$ ts.eps : num 1e-05
$ unzip : chr "/usr/bin/unzip"
$ useFancyQuotes : logi TRUE
$ verbose : logi FALSE
$ warn : num 0
$ warning.length : int 1000
$ width : int 60
int [1:20(1d)] 1 2 3 4 5 1 2 3 4 5 ...
Factor w/ 0 levels:
'data.frame': 0 obs. of 0 variables
language { A + B; list(C, D) }
- attr(*, "srcref")=List of 3
..$ :Class 'srcref' atomic [1:8] 53 12 53 12 12 12 53 53
.. .. ..- attr(*, "srcfile")=Classes 'srcfilecopy', 'srcfile' <environment: 0x33b5738>
..$ :Class 'srcref' atomic [1:8] 53 14 53 16 14 16 53 53
.. .. ..- attr(*, "srcfile")=Classes 'srcfilecopy', 'srcfile' <environment: 0x33b5738>
..$ :Class 'srcref' atomic [1:8] 53 19 53 28 19 28 53 53
.. .. ..- attr(*, "srcfile")=Classes 'srcfilecopy', 'srcfile' <environment: 0x33b5738>
- attr(*, "srcfile")=Classes 'srcfilecopy', 'srcfile' <environment: 0x33b5738>
- attr(*, "wholeSrcref")=Class 'srcref' atomic [1:8] 1 0 53 30 0 30 1 53
.. ..- attr(*, "srcfile")=Classes 'srcfilecopy', 'srcfile' <environment: 0x33b5738>
Loading required package: stats4
Formal class 'mle' [package "stats4"] with 9 slots
..@ call : language mle(minuslogl = ll)
..@ coef : Named num [1:2] 24.99 3.06
.. ..- attr(*, "names")= chr [1:2] "ymax" "xh"
..@ fullcoef : Named num [1:2] 24.99 3.06
.. ..- attr(*, "names")= chr [1:2] "ymax" "xh"
..@ vcov : num [1:2, 1:2] 17.85 -3.72 -3.72 1.07
.. ..- attr(*, "dimnames")=List of 2
.. .. ..$ : chr [1:2] "ymax" "xh"
.. .. ..$ : chr [1:2] "ymax" "xh"
..@ min : num 28.6
..@ details :List of 6
.. ..$ par : Named num [1:2] 24.99 3.06
.. .. ..- attr(*, "names")= chr [1:2] "ymax" "xh"
.. ..$ value : num 28.6
.. ..$ counts : Named int [1:2] 25 18
.. .. ..- attr(*, "names")= chr [1:2] "function" "gradient"
.. ..$ convergence: int 0
.. ..$ message : NULL
.. ..$ hessian : num [1:2, 1:2] 0.203 0.706 0.706 3.388
.. .. ..- attr(*, "dimnames")=List of 2
.. .. .. ..$ : chr [1:2] "ymax" "xh"
.. .. .. ..$ : chr [1:2] "ymax" "xh"
..@ minuslogl:function (ymax = 15, xh = 6)
.. ..- attr(*, "srcref")=Class 'srcref' atomic [1:8] 63 7 64 52 7 52 63 64
.. .. .. ..- attr(*, "srcfile")=Classes 'srcfilecopy', 'srcfile' <environment: 0x33b5738>
..@ nobs : int NA
..@ method : chr "BFGS"
Warning message:
In dpois(y, lambda = ymax/(1 + x/xh), log = TRUE) : NaNs produced
>
> ## and browse them:
> browseEnv()
R objects in .GlobalEnv environment is shown in browser '/usr/bin/firefox'
>
> ## a (simple) function's environment:
> af12 <- approxfun(1:2, 1:2, method = "const")
> browseEnv(envir = environment(af12))
R objects in environment(af12) environment is shown in browser '/usr/bin/firefox'
> # }
>
>
>
>
>
> dev.off()
null device
1
>
|