Supported by Dr. Osamu Ogasawara and  providing providing  . . |
|
Last data update: 2014.03.03 |
Superposes two worm plotsDescriptionSuperposes two worm plots from GAMLSS fitted objects. This is a diagnostic tool for comparing two solutions. Usagewp.twin(obj1, obj2 = NULL, xvar = NULL, xvar.column = 2, n.inter = 16, show.given = FALSE, ylim.worm = 0.5, line = FALSE, cex = 1, col1 = "black", col2 = "orange", warnings = FALSE, ...) Arguments
DetailsThis function is a customized version of the Argument If ValueFor multiple plots the Author(s)Stef van Buuren, using R code of Mikis Stasinopoulos and Bob Rigby ReferencesStasinopoulos D. M. Rigby R.A. (2007) Generalized additive models for location scale and shape (GAMLSS) in R. Journal of Statistical Software, Vol. 23, Issue 7, Dec 2007, http://www.jstatsoft.org/v23/i07. van Buuren and Fredriks M. (2001) Worm plot: simple diagnostic device for modelling growth reference curves. Statistics in Medicine, 20, 1259–1277. van Buuren and Fredriks M. (2007) Worm plot to diagnose fit in quantile regression. Statistical Modelling, 7, 4, 363–376. See Also
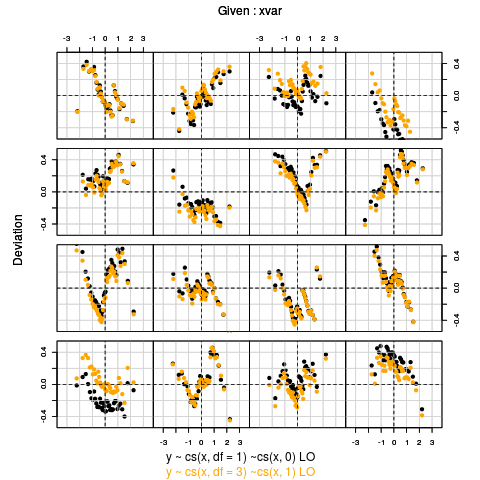
Exampleslibrary(gamlss) data(abdom) a <- gamlss(y~cs(x,df=1),sigma.fo=~cs(x,0),family=LO,data=abdom) b <- gamlss(y~cs(x,df=3),sigma.fo=~cs(x,1),family=LO,data=abdom) coeff1 <- wp.twin(a,b,line=TRUE) coeff1 rm(a,b,coeff1) Results
R version 3.3.1 (2016-06-21) -- "Bug in Your Hair"
Copyright (C) 2016 The R Foundation for Statistical Computing
Platform: x86_64-pc-linux-gnu (64-bit)
R is free software and comes with ABSOLUTELY NO WARRANTY.
You are welcome to redistribute it under certain conditions.
Type 'license()' or 'licence()' for distribution details.
R is a collaborative project with many contributors.
Type 'contributors()' for more information and
'citation()' on how to cite R or R packages in publications.
Type 'demo()' for some demos, 'help()' for on-line help, or
'help.start()' for an HTML browser interface to help.
Type 'q()' to quit R.
> library(AGD)
AGD 0.35 2015-05-27
> png(filename="/home/ddbj/snapshot/RGM3/R_CC/result/AGD/wp.twin.Rd_%03d_medium.png", width=480, height=480)
> ### Name: wp.twin
> ### Title: Superposes two worm plots
> ### Aliases: wp.twin
> ### Keywords: smooth
>
> ### ** Examples
>
> library(gamlss)
Loading required package: splines
Loading required package: gamlss.data
Loading required package: gamlss.dist
Loading required package: MASS
Loading required package: nlme
Loading required package: parallel
********** GAMLSS Version 4.4-0 **********
For more on GAMLSS look at http://www.gamlss.org/
Type gamlssNews() to see new features/changes/bug fixes.
> data(abdom)
> a <- gamlss(y~cs(x,df=1),sigma.fo=~cs(x,0),family=LO,data=abdom)
GAMLSS-RS iteration 1: Global Deviance = 4795.764
GAMLSS-RS iteration 2: Global Deviance = 4797.935
GAMLSS-RS iteration 3: Global Deviance = 4797.948
GAMLSS-RS iteration 4: Global Deviance = 4797.949
GAMLSS-RS iteration 5: Global Deviance = 4797.949
> b <- gamlss(y~cs(x,df=3),sigma.fo=~cs(x,1),family=LO,data=abdom)
GAMLSS-RS iteration 1: Global Deviance = 4781.45
GAMLSS-RS iteration 2: Global Deviance = 4781.096
GAMLSS-RS iteration 3: Global Deviance = 4781.118
GAMLSS-RS iteration 4: Global Deviance = 4781.118
> coeff1 <- wp.twin(a,b,line=TRUE)
> coeff1
$classes
[,1] [,2]
1 12.22 14.64
39 14.64 16.78
77 16.78 18.36
115 18.36 20.07
154 20.07 21.93
192 21.93 23.50
230 23.50 25.07
268 25.07 27.07
306 27.07 28.93
344 28.93 30.64
382 30.64 32.50
420 32.50 34.50
458 34.50 36.21
497 36.21 38.36
535 38.36 40.93
573 40.93 42.50
$coef
[,1] [,2] [,3] [,4]
[1,] -0.003504020 -0.15602304 0.038639888 0.027393705
[2,] 0.026515864 0.28483180 0.006095326 -0.093187380
[3,] -0.077876298 0.05628599 0.061894229 0.009416131
[4,] 0.209698618 -0.02456995 -0.034685801 -0.031058629
[5,] -0.060524040 0.39289973 0.065269080 -0.123323496
[6,] -0.020070631 0.04750037 -0.043512545 -0.042320484
[7,] -0.321110690 -0.10433001 0.168580670 0.056075986
[8,] 0.064037966 -0.05531105 -0.020258555 -0.090437121
[9,] 0.128936714 0.17913097 0.044109902 -0.049297039
[10,] -0.266363310 -0.01890913 0.017866160 -0.019747330
[11,] 0.007205697 -0.11774411 0.137228619 0.021868979
[12,] 0.201188563 0.15077864 -0.052544238 -0.002615670
[13,] 0.007664226 -0.23640794 -0.009871868 0.039585716
[14,] -0.017449540 0.16811355 0.026607674 -0.009545766
[15,] 0.124703184 0.11954586 0.020848864 -0.035485678
[16,] -0.174190902 -0.16016432 -0.081501991 0.003570043
> rm(a,b,coeff1)
>
>
>
>
>
> dev.off()
null device
1
>
|