Supported by Dr. Osamu Ogasawara and  providing providing  . . |
|
Last data update: 2014.03.03 |
Censored Linear Regression Model with Autoregressive ErrorsDescriptionIt fits an univariate left or right censored linear regression model with autoregressive errors under the normal distribution using the SAEM algorithm. It provides estimates and standard errors of the parameters, prediction of future observations and it supports missing values on the dependent variable. It also provides convergence plots when exists at least one censored observation. UsageARCensReg(cc,y,x,p=1,cens='left',x_pred=NULL,miss=NULL, tol=0.0001,show.convergence=TRUE,M=10,perc=0.25,MaxIter=400,pc=0.18) Arguments
DetailsThe initial values are obtained by ignoring censoring and applying maximum likelihood estimation with the censored data simply replaced by their censoring limits. If you want to fit a regression model with autoregressive errors for non-censored data, just set "cc" as a vector of zeros and "cens" as either "right" or "left". Value
Author(s)Fernanda L. Schumacher <fernandalschumacher@gmail.com>, Victor H. Lachos <hlachos@ime.unicamp.br> and Christian E. Galarza <cgalarza88@gmail.com> Maintainer: Fernanda L. Schumacher <fernandalschumacher@gmail.com> ReferencesDelyon, B., Lavielle, M. & Moulines, E. (1999) Convergence of a stochastic approximation version of the EM algorithm. Journal of the Royal Statistical Society, Series B, 39, 1-38. Schumacher, F. L., Lachos, V. H. & Dey, D. K. (2016) Censored regression models with autoregressive errors: A likelihood-based perspective. Submitted. See Also
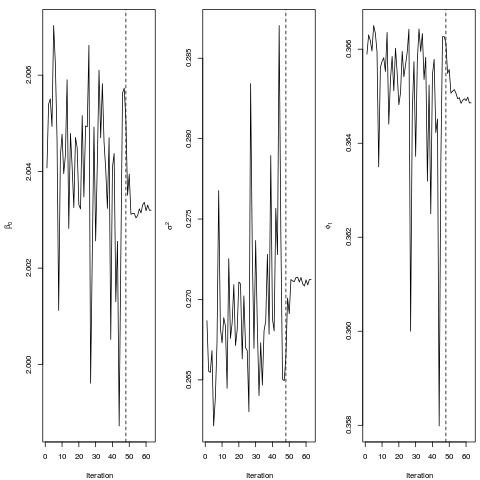
Examples##simple example (p = l = 1) #generating a sample set.seed(23451) phi = .5;p = length(phi) n = 50 sigmae<-.3 beta<-2 pcens = .02 x<-matrix(1,n,1) erro = as.matrix(arima.sim(n,model=list(ar=phi),sd=sqrt(sigmae))) resp<-x%*%beta+erro cte<-as.numeric(quantile(resp,probs=pcens)) cc<-(resp<cte)+0 y<-resp*(1-cc)+cte*cc #fitting the model (quick convergence) fit0 = ARCensReg(cc,y,x,tol=0.0001,pc=.12,M=5) ## Not run: ##another example (p = l = 2) #generating a sample phi = c(.4,-.28) p = length(phi) n = 100 sigmae<-2 beta<-matrix(c(2,1),2,1) pcens = .05 x<-cbind(1,runif(n)) error = as.matrix(arima.sim(n,model=list(ar=phi),sd=sqrt(sigmae))) resp<-x%*%beta+error cte<-as.numeric(quantile(resp,probs=pcens)) cc<-(resp<cte)+0 y<-resp*(1-cc)+cte*cc #fitting the model fit1 = ARCensReg(cc,y,x,p=2,cens="left") #plotting the augmented variable par(mfrow=c(1,1)) plot.ts(fit1$yest,lty='dashed',col=4) lines(y) #simulating missing values miss = sample(1:n,3) y[miss] = NA fit2 = ARCensReg(cc,y,x,p=2,miss=miss,cens="left") #case with no censored values dev.off() fit3 = ARCensReg(rep(0,n),resp,x,p=2) ## End(Not run) Results
R version 3.3.1 (2016-06-21) -- "Bug in Your Hair"
Copyright (C) 2016 The R Foundation for Statistical Computing
Platform: x86_64-pc-linux-gnu (64-bit)
R is free software and comes with ABSOLUTELY NO WARRANTY.
You are welcome to redistribute it under certain conditions.
Type 'license()' or 'licence()' for distribution details.
R is a collaborative project with many contributors.
Type 'contributors()' for more information and
'citation()' on how to cite R or R packages in publications.
Type 'demo()' for some demos, 'help()' for on-line help, or
'help.start()' for an HTML browser interface to help.
Type 'q()' to quit R.
> library(ARCensReg)
> png(filename="/home/ddbj/snapshot/RGM3/R_CC/result/ARCensReg/arcensreg.Rd_%03d_medium.png", width=480, height=480)
> ### Name: ARCensReg
> ### Title: Censored Linear Regression Model with Autoregressive Errors
> ### Aliases: ARCensReg
> ### Keywords: package censored regression autoregressive errors
>
> ### ** Examples
>
> ##simple example (p = l = 1)
> #generating a sample
> set.seed(23451)
> phi = .5;p = length(phi)
> n = 50
> sigmae<-.3
> beta<-2
> pcens = .02
> x<-matrix(1,n,1)
> erro = as.matrix(arima.sim(n,model=list(ar=phi),sd=sqrt(sigmae)))
> resp<-x%*%beta+erro
> cte<-as.numeric(quantile(resp,probs=pcens))
> cc<-(resp<cte)+0
> y<-resp*(1-cc)+cte*cc
>
> #fitting the model (quick convergence)
> fit0 = ARCensReg(cc,y,x,tol=0.0001,pc=.12,M=5)
Call:
ARCensReg(cc = cc, y = y, x = x, tol = 1e-04, M = 5, pc = 0.12)
| | | 0% | |= | 1% | |= | 2% | |== | 2% | |== | 3% | |== | 4% | |=== | 4% | |=== | 5% | |==== | 5% | |==== | 6% | |===== | 6% | |===== | 7% | |===== | 8% | |====== | 8% | |====== | 9% | |======= | 10% | |======== | 11% | |======== | 12% | |========= | 12% | |========= | 13% | |========= | 14% | |========== | 14% | |========== | 15% | |=========== | 15% | |=========== | 16% | |======================================================================| 100%
---------------------------------------------------
Censored Linear Regression Model with AR Errors
---------------------------------------------------
p = 1
---------
Estimates
---------
beta0 sigma2 phi1
2.0032 0.2712 0.3649
s.e. 0.1147 0.0548 0.1305
------------------------
Model selection criteria
------------------------
Loglik AIC BIC AICcorr
Value -38.816 83.631 89.367 84.153
-------
Details
-------
Type of censoring = left
Number of missing values = 0
Convergence reached? = TRUE
Iterations = 63 / 400
MC sample = 5
Cut point = 0.12
Processing time = 4.151549 secs
>
> ## Not run:
> ##D
> ##D ##another example (p = l = 2)
> ##D #generating a sample
> ##D phi = c(.4,-.28)
> ##D p = length(phi)
> ##D n = 100
> ##D sigmae<-2
> ##D beta<-matrix(c(2,1),2,1)
> ##D pcens = .05
> ##D x<-cbind(1,runif(n))
> ##D error = as.matrix(arima.sim(n,model=list(ar=phi),sd=sqrt(sigmae)))
> ##D resp<-x%*%beta+error
> ##D cte<-as.numeric(quantile(resp,probs=pcens))
> ##D cc<-(resp<cte)+0
> ##D y<-resp*(1-cc)+cte*cc
> ##D
> ##D #fitting the model
> ##D fit1 = ARCensReg(cc,y,x,p=2,cens="left")
> ##D
> ##D #plotting the augmented variable
> ##D par(mfrow=c(1,1))
> ##D plot.ts(fit1$yest,lty='dashed',col=4)
> ##D lines(y)
> ##D
> ##D #simulating missing values
> ##D miss = sample(1:n,3)
> ##D y[miss] = NA
> ##D fit2 = ARCensReg(cc,y,x,p=2,miss=miss,cens="left")
> ##D
> ##D #case with no censored values
> ##D dev.off()
> ##D fit3 = ARCensReg(rep(0,n),resp,x,p=2)
> ## End(Not run)
>
>
>
>
>
> dev.off()
null device
1
>
|