Supported by Dr. Osamu Ogasawara and  providing providing  . . |
|
Last data update: 2014.03.03 |
Independence Chain Metropolis-Hastings Algorithm using an Adaptive Mixture of Student-t Distributions as the Candidate DensityDescriptionPerforms independence chain Metropolis-Hastings (M-H) sampling using an adaptive mixture of Student-t distributions as the candidate density UsageAdMitMH(N = 1e5, KERNEL, mit = list(), ...) Arguments
DetailsThe argument
where H (>=1) is the number of components and
d (>=1) is the dimension of the first argument in ValueA list with the following components:
NoteFurther details and examples of the R package Further information on the Metropolis-Hastings algorithm can be found in Chib and Greenberg (1995) and Koop (2003). Please cite the package in publications. Use Author(s)David Ardia ReferencesArdia, D., Hoogerheide, L.F., van Dijk, H.K. (2009a). AdMit: Adaptive Mixture of Student-t Distributions. The R Journal 1(1), pp.25–30. http://journal.r-project.org/2009-1/ Ardia, D., Hoogerheide, L.F., van Dijk, H.K. (2009b). Adaptive Mixture of Student-t Distributions as a Flexible Candidate Distribution for Efficient Simulation: The R Package AdMit. Journal of Statistical Software 29(3), pp.1–32. http://www.jstatsoft.org/v29/i03/ Chib, S., Greenberg, E. (1995). Understanding the Metropolis-Hasting Algorithm. The American Statistician 49(4), pp.327–335. Koop, G. (2003). Bayesian Econometrics. Wiley-Interscience (London, UK). ISBN: 0470845678. See Also
Examples
## NB : Low number of draws for speedup. Consider using more draws!
## Gelman and Meng (1991) kernel function
GelmanMeng <- function(x, A = 1, B = 0, C1 = 3, C2 = 3, log = TRUE)
{
if (is.vector(x))
x <- matrix(x, nrow = 1)
r <- -.5 * (A * x[,1]^2 * x[,2]^2 + x[,1]^2 + x[,2]^2
- 2 * B * x[,1] * x[,2] - 2 * C1 * x[,1] - 2 * C2 * x[,2])
if (!log)
r <- exp(r)
as.vector(r)
}
## Run the AdMit function to fit the mixture approximation
set.seed(1234)
outAdMit <- AdMit(KERNEL = GelmanMeng,
mu0 = c(0.0, 0.1), control = list(Ns = 1e4))
## Run M-H using the mixture approximation as the candidate density
outAdMitMH <- AdMitMH(N = 1e4, KERNEL = GelmanMeng, mit = outAdMit$mit)
options(digits = 4, max.print = 40)
print(outAdMitMH)
## Use functions provided by the package coda to obtain
## quantities of interest for the density whose kernel is 'GelmanMeng'
library("coda")
draws <- as.mcmc(outAdMitMH$draws)
draws <- window(draws, start = 1001)
colnames(draws) <- c("X1", "X2")
summary(draws)
summary(draws)$stat[,3]^2 / summary(draws)$stat[,4]^2 ## RNE
plot(draws)
Results
R version 3.3.1 (2016-06-21) -- "Bug in Your Hair"
Copyright (C) 2016 The R Foundation for Statistical Computing
Platform: x86_64-pc-linux-gnu (64-bit)
R is free software and comes with ABSOLUTELY NO WARRANTY.
You are welcome to redistribute it under certain conditions.
Type 'license()' or 'licence()' for distribution details.
R is a collaborative project with many contributors.
Type 'contributors()' for more information and
'citation()' on how to cite R or R packages in publications.
Type 'demo()' for some demos, 'help()' for on-line help, or
'help.start()' for an HTML browser interface to help.
Type 'q()' to quit R.
> library(AdMit)
Loading required package: mvtnorm
> png(filename="/home/ddbj/snapshot/RGM3/R_CC/result/AdMit/AdMitMH.Rd_%03d_medium.png", width=480, height=480)
> ### Name: AdMitMH
> ### Title: Independence Chain Metropolis-Hastings Algorithm using an
> ### Adaptive Mixture of Student-t Distributions as the Candidate Density
> ### Aliases: AdMitMH
> ### Keywords: htest
>
> ### ** Examples
>
> ## NB : Low number of draws for speedup. Consider using more draws!
> ## Gelman and Meng (1991) kernel function
> GelmanMeng <- function(x, A = 1, B = 0, C1 = 3, C2 = 3, log = TRUE)
+ {
+ if (is.vector(x))
+ x <- matrix(x, nrow = 1)
+ r <- -.5 * (A * x[,1]^2 * x[,2]^2 + x[,1]^2 + x[,2]^2
+ - 2 * B * x[,1] * x[,2] - 2 * C1 * x[,1] - 2 * C2 * x[,2])
+ if (!log)
+ r <- exp(r)
+ as.vector(r)
+ }
>
> ## Run the AdMit function to fit the mixture approximation
> set.seed(1234)
> outAdMit <- AdMit(KERNEL = GelmanMeng,
+ mu0 = c(0.0, 0.1), control = list(Ns = 1e4))
>
> ## Run M-H using the mixture approximation as the candidate density
> outAdMitMH <- AdMitMH(N = 1e4, KERNEL = GelmanMeng, mit = outAdMit$mit)
> options(digits = 4, max.print = 40)
> print(outAdMitMH)
$draws
k1 k2
1 1.584e+00 -3.8213044
2 1.584e+00 -3.8213044
3 2.148e+00 0.3020074
4 2.148e+00 0.3020074
5 2.148e+00 0.3020074
6 2.148e+00 0.3020074
7 2.451e+00 0.2128177
8 2.451e+00 0.2128177
9 2.451e+00 0.2128177
10 2.451e+00 0.2128177
11 7.927e-01 1.4841330
12 7.927e-01 1.4841330
13 9.880e-01 0.6008210
14 9.880e-01 0.6008210
15 9.292e-01 0.5884448
16 9.292e-01 0.5884448
17 7.471e-01 1.1384726
18 2.444e+00 0.2993253
19 3.022e+00 0.0115478
20 3.022e+00 0.0115478
[ reached getOption("max.print") -- omitted 9980 rows ]
$accept
[1] 0.5328
>
> ## Use functions provided by the package coda to obtain
> ## quantities of interest for the density whose kernel is 'GelmanMeng'
> library("coda")
> draws <- as.mcmc(outAdMitMH$draws)
> draws <- window(draws, start = 1001)
> colnames(draws) <- c("X1", "X2")
> summary(draws)
Iterations = 1001:10000
Thinning interval = 1
Number of chains = 1
Sample size per chain = 9000
1. Empirical mean and standard deviation for each variable,
plus standard error of the mean:
Mean SD Naive SE Time-series SE
X1 1.46 1.24 0.0131 0.0219
X2 1.47 1.25 0.0132 0.0217
2. Quantiles for each variable:
2.5% 25% 50% 75% 97.5%
X1 -0.189 0.425 1.19 2.36 4.17
X2 -0.185 0.422 1.15 2.36 4.22
> summary(draws)$stat[,3]^2 / summary(draws)$stat[,4]^2 ## RNE
X1 X2
0.3568 0.3673
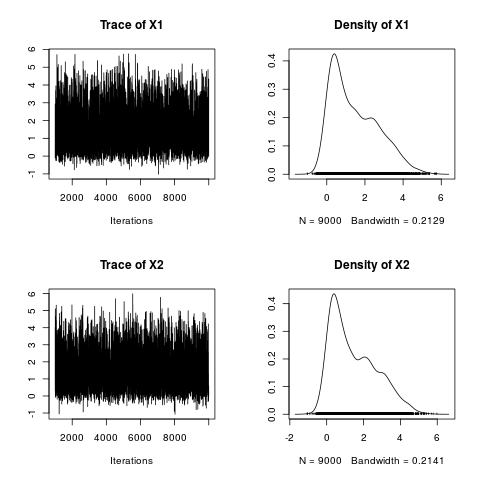
> plot(draws)
>
>
>
>
>
> dev.off()
null device
1
>
|