Supported by Dr. Osamu Ogasawara and  providing providing  . . |
|
Last data update: 2014.03.03 |
Cluster Sequences By Distance or SequenceDescriptionGroups the sequences represented by a distance matrix into clusters of similarity. Usage
IdClusters(myDistMatrix = NULL,
method = "UPGMA",
cutoff = -Inf,
showPlot = FALSE,
asDendrogram = FALSE,
myXStringSet = NULL,
model = MODELS,
processors = 1,
verbose = TRUE)
Arguments
Details
Ultrametric methods: The method Additive methods: Sequence-only method: Multiple cutoffs may be provided if they are in increasing or decreasing order. If ValueIf Author(s)Erik Wright DECIPHER@cae.wisc.edu ReferencesFelsenstein, J. (1981) Evolutionary trees from DNA sequences: a maximum likelihood approach. Journal of Molecular Evolution, 17(6), 368-376. Ghodsi, M., Liu, B., & Pop, M. (2011) DNACLUST. BMC Bioinformatics, 12(1), 271. doi:10.1186/1471-2105-12-271. Saitou, N. and Nei, M. (1987) The neighbor-joining method: a new method for reconstructing phylogenetic trees. Molecular Biology and Evolution, 4(4), 406-425. See Also
Examples
# using the matrix from the original paper by Saitou and Nei
m <- matrix(0,8,8)
m[2:8,1] <- c(7, 8, 11, 13, 16, 13, 17)
m[3:8,2] <- c(5, 8, 10, 13, 10, 14)
m[4:8,3] <- c(5, 7, 10, 7, 11)
m[5:8,4] <- c(8, 11, 8, 12)
m[6:8,5] <- c(5, 6, 10)
m[7:8,6] <- c(9, 13)
m[8,7] <- c(8)
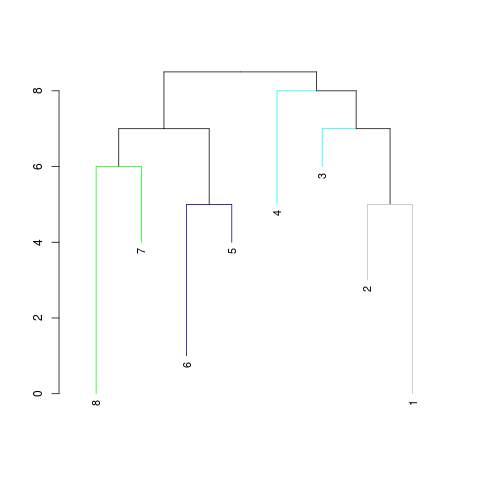
# returns an object of class "dendrogram"
myClusters <- IdClusters(m, cutoff=10, method="NJ", showPlot=TRUE, asDendrogram=TRUE)
# example of specifying multiple cutoffs
IdClusters(m, cutoff=c(2,6,10,20)) # returns a data frame
# example of 'inexact' clustering
fas <- system.file("extdata", "50S_ribosomal_protein_L2.fas", package="DECIPHER")
dna <- readDNAStringSet(fas)
IdClusters(myXStringSet=dna, method="inexact", cutoff=0.05)
Results
R version 3.3.1 (2016-06-21) -- "Bug in Your Hair"
Copyright (C) 2016 The R Foundation for Statistical Computing
Platform: x86_64-pc-linux-gnu (64-bit)
R is free software and comes with ABSOLUTELY NO WARRANTY.
You are welcome to redistribute it under certain conditions.
Type 'license()' or 'licence()' for distribution details.
R is a collaborative project with many contributors.
Type 'contributors()' for more information and
'citation()' on how to cite R or R packages in publications.
Type 'demo()' for some demos, 'help()' for on-line help, or
'help.start()' for an HTML browser interface to help.
Type 'q()' to quit R.
> library(DECIPHER)
Loading required package: Biostrings
Loading required package: BiocGenerics
Loading required package: parallel
Attaching package: 'BiocGenerics'
The following objects are masked from 'package:parallel':
clusterApply, clusterApplyLB, clusterCall, clusterEvalQ,
clusterExport, clusterMap, parApply, parCapply, parLapply,
parLapplyLB, parRapply, parSapply, parSapplyLB
The following objects are masked from 'package:stats':
IQR, mad, xtabs
The following objects are masked from 'package:base':
Filter, Find, Map, Position, Reduce, anyDuplicated, append,
as.data.frame, cbind, colnames, do.call, duplicated, eval, evalq,
get, grep, grepl, intersect, is.unsorted, lapply, lengths, mapply,
match, mget, order, paste, pmax, pmax.int, pmin, pmin.int, rank,
rbind, rownames, sapply, setdiff, sort, table, tapply, union,
unique, unsplit
Loading required package: S4Vectors
Loading required package: stats4
Attaching package: 'S4Vectors'
The following objects are masked from 'package:base':
colMeans, colSums, expand.grid, rowMeans, rowSums
Loading required package: IRanges
Loading required package: XVector
Loading required package: RSQLite
Loading required package: DBI
> png(filename="/home/ddbj/snapshot/RGM3/R_BC/result/DECIPHER/IdClusters.Rd_%03d_medium.png", width=480, height=480)
> ### Name: IdClusters
> ### Title: Cluster Sequences By Distance or Sequence
> ### Aliases: IdClusters
>
> ### ** Examples
>
> # using the matrix from the original paper by Saitou and Nei
> m <- matrix(0,8,8)
> m[2:8,1] <- c(7, 8, 11, 13, 16, 13, 17)
> m[3:8,2] <- c(5, 8, 10, 13, 10, 14)
> m[4:8,3] <- c(5, 7, 10, 7, 11)
> m[5:8,4] <- c(8, 11, 8, 12)
> m[6:8,5] <- c(5, 6, 10)
> m[7:8,6] <- c(9, 13)
> m[8,7] <- c(8)
>
> # returns an object of class "dendrogram"
> myClusters <- IdClusters(m, cutoff=10, method="NJ", showPlot=TRUE, asDendrogram=TRUE)
| | | 0% | |================== | 25% | |================================ | 46% | |============================================= | 64% | |======================================================= | 78% | |============================================================== | 89% | |=================================================================== | 96% | |======================================================================| 100%
Time difference of 0.06 secs
>
> # example of specifying multiple cutoffs
> IdClusters(m, cutoff=c(2,6,10,20)) # returns a data frame
| | | 0% | |================== | 25% | |================================ | 46% | |============================================= | 64% | |======================================================= | 78% | |============================================================== | 89% | |=================================================================== | 96% | |======================================================================| 100%
Time difference of 0.01 secs
cluster2UPGMA cluster6UPGMA cluster10UPGMA cluster20UPGMA
1 7 5 2 1
2 5 3 2 1
3 4 2 2 1
4 3 2 2 1
5 2 1 1 1
6 1 1 1 1
7 6 4 1 1
8 8 6 3 1
>
> # example of 'inexact' clustering
> fas <- system.file("extdata", "50S_ribosomal_protein_L2.fas", package="DECIPHER")
> dna <- readDNAStringSet(fas)
> IdClusters(myXStringSet=dna, method="inexact", cutoff=0.05)
| | | 0% | |= | 1% | |= | 2% | |== | 3% | |=== | 4% | |==== | 5% | |==== | 6% | |===== | 7% | |====== | 8% | |====== | 9% | |======= | 10% | |======== | 11% | |======== | 12% | |========= | 13% | |========== | 14% | |========== | 15% | |=========== | 16% | |============ | 17% | |============= | 18% | |============= | 19% | |============== | 20% | |=============== | 21% | |=============== | 22% | |================ | 23% | |================= | 24% | |================== | 25% | |================== | 26% | |=================== | 27% | |==================== | 28% | |==================== | 29% | |===================== | 30% | |====================== | 31% | |====================== | 32% | |======================= | 33% | |======================== | 34% | |======================== | 35% | |========================= | 36% | |========================== | 37% | |=========================== | 38% | |=========================== | 39% | |============================ | 40% | |============================= | 41% | |============================= | 42% | |============================== | 43% | |=============================== | 44% | |================================ | 45% | |================================ | 46% | |================================= | 47% | |================================== | 48% | |================================== | 49% | |=================================== | 50% | |==================================== | 51% | |==================================== | 52% | |===================================== | 53% | |====================================== | 54% | |====================================== | 55% | |======================================= | 56% | |======================================== | 57% | |========================================= | 58% | |========================================= | 59% | |========================================== | 60% | |=========================================== | 61% | |=========================================== | 62% | |============================================ | 63% | |============================================= | 64% | |============================================== | 65% | |============================================== | 66% | |=============================================== | 67% | |================================================ | 68% | |================================================ | 69% | |================================================= | 70% | |================================================== | 71% | |================================================== | 72% | |=================================================== | 73% | |==================================================== | 74% | |==================================================== | 75% | |===================================================== | 76% | |====================================================== | 77% | |======================================================= | 78% | |======================================================= | 79% | |======================================================== | 80% | |========================================================= | 81% | |========================================================= | 82% | |========================================================== | 83% | |=========================================================== | 84% | |============================================================ | 85% | |============================================================ | 86% | |============================================================= | 87% | |============================================================== | 88% | |============================================================== | 89% | |=============================================================== | 90% | |================================================================ | 91% | |================================================================ | 92% | |================================================================= | 93% | |================================================================== | 94% | |================================================================== | 95% | |=================================================================== | 96% | |==================================================================== | 97% | |===================================================================== | 98% | |===================================================================== | 99% | |======================================================================| 100%
Time difference of 3.25 secs
cluster
Rickettsia prowazekii str. Dachau 102
Porphyromonas gingivalis W83 90
Porphyromonas gingivalis TDC60 90
Porphyromonas gingivalis ATCC 33277 90
Pasteurella multocida 671/90 103
Pasteurella multocida 36950 103
Xanthomonas campestris pv. campestris 79
Lactobacillus plantarum subsp. plantarum P-8 27
Lactobacillus plantarum ZJ316 27
Lactobacillus plantarum subsp. plantarum NC8 27
Lactobacillus plantarum subsp. plantarum ATCC 14917 27
Lactobacillus plantarum WCFS1 27
Xanthomonas citri pv. mangiferaeindicae LMG 941 80
Xanthomonas axonopodis pv. punicae str. LMG 859 80
Xanthomonas citri subsp. citri Aw12879 80
Xanthomonas vesicatoria ATCC 35937 80
Xanthomonas fuscans subsp. aurantifolii str. ICPB 11122 80
Xanthomonas fuscans subsp. aurantifolii str. ICPB 10535 80
Xanthomonas campestris pv. vesicatoria str. 85-10 80
Xanthomonas axonopodis pv. citrumelo F1 80
Xanthomonas oryzae pv. oryzicola BLS256 81
Xanthomonas campestris pv. raphani 756C 79
Bacillus halodurans C-125 56
Corynebacterium glutamicum SCgG2 20
Corynebacterium glutamicum K051 20
Synechocystis sp. PCC 6803 substr. PCC-N 57
Vibrio parahaemolyticus O1:K33 str. CDC_K4557 91
Vibrio parahaemolyticus BB22OP 91
Vibrio parahaemolyticus 10329 91
Streptococcus pyogenes SSI-1 115
Lactococcus lactis subsp. lactis IO-1 58
Lactococcus lactis subsp. lactis bv. diacetylactis str. LD61 58
Lactococcus lactis subsp. cremoris CNCM I-1631 58
Lactococcus lactis subsp. lactis KF147 58
Clostridium perfringens F262 41
Clostridium perfringens D str. JGS1721 41
Clostridium perfringens SM101 41
Mycoplasma pneumoniae M129-B7 4
Streptomyces avermitilis MA-4680 32
Treponema pallidum subsp. pallidum str. Nichols 104
Helicobacter pylori X47-2AL 59
Helicobacter pylori G27 60
Helicobacter pylori XZ274 61
Helicobacter pylori Shi417 61
Helicobacter pylori UM032 61
Helicobacter pylori Aklavik86 61
Helicobacter pylori 83 61
Helicobacter pylori Hp M3 62
Helicobacter pylori Hp P-15b 60
Helicobacter pylori Hp P-4d 62
Helicobacter pylori Hp P-62 62
Helicobacter pylori Hp P-30 60
Helicobacter pylori Hp P-3 62
Helicobacter pylori Hp H-23 62
Helicobacter pylori Hp H-19 62
Helicobacter pylori Hp H-18 62
Helicobacter pylori Hp H-11 59
Helicobacter pylori Hp A-16 62
Helicobacter pylori Hp H-44 62
Helicobacter pylori Hp H-41 62
Helicobacter pylori Hp H-27 59
Helicobacter pylori CPY6311 61
Helicobacter pylori CPY1313 61
Helicobacter pylori NQ4216 59
Helicobacter pylori Hp H-16 62
Helicobacter pylori Hp H-9 59
Helicobacter pylori Hp H-30 62
Helicobacter pylori Hp P-11 62
Helicobacter pylori CPY1962 59
Helicobacter pylori Hp H-4 62
Helicobacter pylori Hp H-5b 62
Helicobacter pylori CPY1124 59
Helicobacter pylori Hp H-42 62
Helicobacter pylori Hp H-36 62
Helicobacter pylori Hp H-10 62
Helicobacter pylori Hp P-74 59
Helicobacter pylori NQ4044 60
Helicobacter pylori Hp A-26 60
Helicobacter pylori CPY6081 59
Helicobacter pylori CPY3281 61
Helicobacter pylori NQ4200 59
Helicobacter pylori Hp A-8 62
Helicobacter pylori NQ4053 62
Helicobacter pylori Hp P-26 62
Helicobacter pylori CPY6261 59
Helicobacter pylori B128 62
Helicobacter pylori 98-10 61
Helicobacter pylori Hp A-6 62
Helicobacter pylori ELS37 59
Helicobacter pylori Puno135 59
Helicobacter pylori Puno120 59
Helicobacter pylori Gambia94/24 62
Helicobacter pylori 2017 62
Helicobacter pylori Sat464 59
Helicobacter pylori 908 62
Helicobacter pylori v225d 59
Helicobacter pylori J99 60
Helicobacter pylori P12 59
Helicobacter pylori F57 59
Helicobacter pylori F32 61
Helicobacter pylori F16 63
Helicobacter pylori Rif2 59
Helicobacter pylori PeCan18 64
Helicobacter pylori Hp P-13b 62
Helicobacter pylori Hp P-16 60
Helicobacter pylori Hp H-43 62
Helicobacter pylori Hp A-9 59
Hel
|