Supported by Dr. Osamu Ogasawara and  providing providing  . . |
|
Last data update: 2014.03.03 |
Calculate Maximum Likelihood Estimates for a 2D GQD Model.Description
where
and
Usage
BiGQD.mle(X, time, mesh=10, theta, control=NULL, method='Nelder-Mead', RK.order=4,
exclude=NULL, Tag=NA, Dtype='Saddlepoint', rtf= runif(2,-1,1), wrt=FALSE,
print.output=TRUE)
Arguments
Value
Syntactical jargonSynt. [1]: The coefficients of the 2D GQD may be parameterized using the reserved variable
Synt. [2]: Due to syntactical differences between R and C++ special functions have to be used when terms that depend on
Here sqrt(t)*cos(3*pi*t) constitutes the product of two terms that cannot be written i.t.o. a single Synt. [3]: Similarly, the ^ - operator is not overloaded in C++. Instead the
WarningWarning [1]: The parameter Warning [2]: Note that minus the likelihood is minimized, as such the Author(s)Etienne A.D. Pienaar: etiennead@gmail.com ReferencesUpdates available on GitHub at https://github.com/eta21. Daniels, H.E. 1954 Saddlepoint approximations in statistics. Ann. Math. Stat., 25:631–650. Eddelbuettel, D. and Romain, F. 2011 Rcpp: Seamless R and C++ integration. Journal of Statistical Software, 40(8):1–18,. URL http://www.jstatsoft.org/v40/i08/. Eddelbuettel, D. 2013 Seamless R and C++ Integration with Rcpp. New York: Springer. ISBN 978-1-4614-6867-7. Eddelbuettel, D. and Sanderson, C. 2014 Rcpparmadillo: Accelerating R with high-performance C++ linear algebra. Computational Statistics and Data Analysis, 71:1054–1063. URL http://dx.doi.org/10.1016/j.csda.2013.02.005. Feagin, T. 2007 A tenth-order Runge-Kutta method with error estimate. In Proceedings of the IAENG Conf. on Scientifc Computing. Varughese, M.M. 2013 Parameter estimation for multivariate diffusion systems. Comput. Stat. Data An., 57:417–428. See Also
Examples
#===============================================================================
# This example simulates a bivariate time homogeneous diffusion and shows how
# to conduct inference using BiGQD.mle(). We fit two competing models and then
# use the output to select a winner.
#-------------------------------------------------------------------------------
data(SDEsim2)
data(SDEsim2)
attach(SDEsim2)
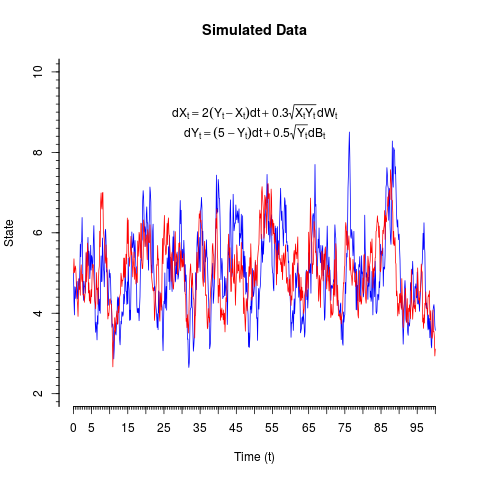
# Have a look at the time series:
plot(Xt~time,type='l',col='blue',ylim=c(2,10),main='Simulated Data',xlab='Time (t)',ylab='State',
axes=FALSE)
lines(Yt~time,col='red')
expr1=expression(dX[t]==2(Y[t]-X[t])*dt+0.3*sqrt(X[t]*Y[t])*dW[t])
expr2=expression(dY[t]==(5-Y[t])*dt+0.5*sqrt(Y[t])*dB[t])
text(50,9,expr1)
text(50,8.5,expr2)
axis(1,seq(0,100,5))
axis(1,seq(0,100,5/10),tcl=-0.2,labels=NA)
axis(2,seq(0,20,2))
axis(2,seq(0,20,2/10),tcl=-0.2,labels=NA)
#------------------------------------------------------------------------------
# Define the coefficients of a proposed model
#------------------------------------------------------------------------------
GQD.remove()
a00 <- function(t){theta[1]*theta[2]}
a10 <- function(t){-theta[1]}
c00 <- function(t){theta[3]*theta[3]}
b00 <- function(t){theta[4]}
b01 <- function(t){-theta[5]}
f00 <- function(t){theta[6]*theta[6]}
theta.start <- c(3,3,3,3,3,3)
X <- cbind(Xt,Yt)
# Calculate MLEs
m1=BiGQD.mle(X,time,10,theta.start)
#------------------------------------------------------------------------------
# Remove old coefficients and define the coefficients of a new model
#------------------------------------------------------------------------------
GQD.remove()
a10 <- function(t){-theta[1]}
a01 <- function(t){theta[1]*theta[2]}
c11 <- function(t){theta[3]*theta[3]}
b00 <- function(t){theta[4]*theta[5]}
b01 <- function(t){-theta[4]}
f01 <- function(t){theta[6]*theta[6]}
theta.start <- c(3,3,3,3,3,3)
# Calculate MLEs
m2=BiGQD.mle(X,time,10,theta.start)
# Compare estimates:
GQD.estimates(m1)
GQD.estimates(m2)
#------------------------------------------------------------------------------
# Compare the two models
#------------------------------------------------------------------------------
GQD.aic(list(m1,m2))
#===============================================================================
Results
R version 3.3.1 (2016-06-21) -- "Bug in Your Hair"
Copyright (C) 2016 The R Foundation for Statistical Computing
Platform: x86_64-pc-linux-gnu (64-bit)
R is free software and comes with ABSOLUTELY NO WARRANTY.
You are welcome to redistribute it under certain conditions.
Type 'license()' or 'licence()' for distribution details.
R is a collaborative project with many contributors.
Type 'contributors()' for more information and
'citation()' on how to cite R or R packages in publications.
Type 'demo()' for some demos, 'help()' for on-line help, or
'help.start()' for an HTML browser interface to help.
Type 'q()' to quit R.
> library(DiffusionRgqd)
> png(filename="/home/ddbj/snapshot/RGM3/R_CC/result/DiffusionRgqd/BiGQD.mle.Rd_%03d_medium.png", width=480, height=480)
> ### Name: BiGQD.mle
> ### Title: Calculate Maximum Likelihood Estimates for a 2D GQD Model.
> ### Aliases: BiGQD.mle
> ### Keywords: syntax C++ maximum likelihood
>
> ### ** Examples
>
> ## No test:
> #===============================================================================
> # This example simulates a bivariate time homogeneous diffusion and shows how
> # to conduct inference using BiGQD.mle(). We fit two competing models and then
> # use the output to select a winner.
> #-------------------------------------------------------------------------------
>
> data(SDEsim2)
> data(SDEsim2)
> attach(SDEsim2)
> # Have a look at the time series:
> plot(Xt~time,type='l',col='blue',ylim=c(2,10),main='Simulated Data',xlab='Time (t)',ylab='State',
+ axes=FALSE)
> lines(Yt~time,col='red')
> expr1=expression(dX[t]==2(Y[t]-X[t])*dt+0.3*sqrt(X[t]*Y[t])*dW[t])
> expr2=expression(dY[t]==(5-Y[t])*dt+0.5*sqrt(Y[t])*dB[t])
> text(50,9,expr1)
> text(50,8.5,expr2)
> axis(1,seq(0,100,5))
> axis(1,seq(0,100,5/10),tcl=-0.2,labels=NA)
> axis(2,seq(0,20,2))
> axis(2,seq(0,20,2/10),tcl=-0.2,labels=NA)
>
> #------------------------------------------------------------------------------
> # Define the coefficients of a proposed model
> #------------------------------------------------------------------------------
> GQD.remove()
[1] "Removed : NA "
> a00 <- function(t){theta[1]*theta[2]}
> a10 <- function(t){-theta[1]}
> c00 <- function(t){theta[3]*theta[3]}
>
> b00 <- function(t){theta[4]}
> b01 <- function(t){-theta[5]}
> f00 <- function(t){theta[6]*theta[6]}
>
> theta.start <- c(3,3,3,3,3,3)
> X <- cbind(Xt,Yt)
>
> # Calculate MLEs
> m1=BiGQD.mle(X,time,10,theta.start)
Compiling C++ code. Please wait.
================================================================
GENERALIZED LINEAR DIFFUSON
================================================================
_____________________ Drift Coefficients _______________________
a00 : theta[1]*theta[2]
a10 : -theta[1]
... ... ... ... ... ... ... ... ... ... ...
b00 : theta[4]
b01 : -theta[5]
___________________ Diffusion Coefficients _____________________
c00 : theta[3]*theta[3]
... ... ... ... ... ... ... ... ... ... ...
... ... ... ... ... ... ... ... ... ... ...
... ... ... ... ... ... ... ... ... ... ...
f00 : theta[6]*theta[6]
__________________________ Model Info __________________________
Time Homogeneous : Yes
Data Resolution : Homogeneous: dt=0.125
# Removed Transits. : None
Density approx. : 2nd Ord. Truncation, Bivariate Normal
Elapsed time : 00:00:01
... ... ... ... ... ... ... ... ... ... ...
dim(theta) : 6
----------------------------------------------------------------
>
> #------------------------------------------------------------------------------
> # Remove old coefficients and define the coefficients of a new model
> #------------------------------------------------------------------------------
> GQD.remove()
[1] "Removed : a00 a10 b00 b01 c00 f00"
> a10 <- function(t){-theta[1]}
> a01 <- function(t){theta[1]*theta[2]}
> c11 <- function(t){theta[3]*theta[3]}
>
> b00 <- function(t){theta[4]*theta[5]}
> b01 <- function(t){-theta[4]}
> f01 <- function(t){theta[6]*theta[6]}
>
> theta.start <- c(3,3,3,3,3,3)
>
> # Calculate MLEs
> m2=BiGQD.mle(X,time,10,theta.start)
Compiling C++ code. Please wait.
================================================================
GENERALIZED QUADRATIC DIFFUSON
================================================================
_____________________ Drift Coefficients _______________________
a10 : -theta[1]
a01 : theta[1]*theta[2]
... ... ... ... ... ... ... ... ... ... ...
b00 : theta[4]*theta[5]
b01 : -theta[4]
___________________ Diffusion Coefficients _____________________
c11 : theta[3]*theta[3]
... ... ... ... ... ... ... ... ... ... ...
... ... ... ... ... ... ... ... ... ... ...
... ... ... ... ... ... ... ... ... ... ...
f01 : theta[6]*theta[6]
__________________________ Model Info __________________________
Time Homogeneous : Yes
Data Resolution : Homogeneous: dt=0.125
# Removed Transits. : None
Density approx. : 4th Ord. Truncation, Bivariate-Saddlepoint
Elapsed time : 00:00:07
... ... ... ... ... ... ... ... ... ... ...
dim(theta) : 6
----------------------------------------------------------------
>
> # Compare estimates:
> GQD.estimates(m1)
Estimate Lower_95 Upper_95
theta[1] 1.073 0.749 1.397
theta[2] 5.023 4.735 5.311
theta[3] 1.495 1.418 1.573
theta[4] 5.377 3.714 7.040
theta[5] 1.075 0.748 1.403
theta[6] 1.224 1.160 1.287
> GQD.estimates(m2)
Estimate Lower_95 Upper_95
theta[1] 5.639 NaN NaN
theta[2] 0.994 0.978 1.010
theta[3] 0.421 NaN NaN
theta[4] 3.077 2.361 3.792
theta[5] 5.079 4.968 5.189
theta[6] 0.660 0.601 0.719
Warning message:
In sqrt(diag(solve(-x$opt$hessian))) : NaNs produced
>
> #------------------------------------------------------------------------------
> # Compare the two models
> #------------------------------------------------------------------------------
>
> GQD.aic(list(m1,m2))
Convergence p min.likelihood AIC BIC N
Model 1 0 6 910.9934 [=] 1833.987 [=] 1862.102 801
Model 2 0 6 1047.7193 2107.4385 2135.5537 801
>
>
> #===============================================================================
> ## End(No test)
>
>
>
>
>
> dev.off()
null device
1
>
|