Supported by Dr. Osamu Ogasawara and  providing providing  . . |
|
Last data update: 2014.03.03 |
GS-Estimates for multivariate regression with bootstrap confidence intervalsDescriptionComputes GS-estimates for multivariate regression together with standard errors, confidence intervals and p-values based on the Fast and Robust Bootstrap. Usage## S3 method for class 'formula' FRBmultiregGS(formula, data=NULL, ...) ## Default S3 method: FRBmultiregGS(X, Y, int = TRUE, R = 999, bdp = 0.5, conf = 0.95, control=GScontrol(...), na.action=na.omit, ...) Arguments
Details Generalized S-estimators are defined by minimizing the determinant of a robust estimator of the scatter matrix of
the differences of the residuals (Roelant et al. 2009). Hence, this procedure is intercept free and only gives an estimate for the slope matrix. To estimate
the intercept, we use the M-type estimator of location of Lopuhaa (1992) on the residuals with the residual scatter matrix
estimate of the residuals as a preliminary estimate. This computation is carried out by a call to The Fast and Robust Bootstrap (Salibian-Barrera and Zamar 2002) is used to calculate so-called
basic bootstrap confidence intervals and bias corrected and accelerated (BCa)
confidence intervals (Davison and Hinkley 1997, p.194 and p.204 respectively).
Apart from the intervals with the requested confidence level, the function also returns p-values for each coefficient
corresponding to the hypothesis that the actual coefficient is zero. The p-values are computed as
1 minus the smallest level for which the confidence intervals would include zero. Both BCa and basic bootstrap p-values in this sense are given.
The bootstrap calculation is carried out by a call to Note: Bootstrap samples which contain too few distinct observations with positive weights are discarded
(a warning is given if this happens). The number of samples actually used is returned via In the The returned object inherits from class ValueAn object of class
Author(s)Ella Roelant, Stefan Van Aelst and Gert Willems References
See Also
Examples
data(schooldata)
school.x <- data.matrix(schooldata[,1:5])
school.y <- data.matrix(schooldata[,6:8])
#computes 25% breakdown point GS-estimate and 80% confidence intervals
#based on 99 bootstrap samples:
GSres <- FRBmultiregGS(school.x, school.y, R=99, bdp = 0.25, conf = 0.8,nsamp=50)
#or using the formula interface
## Not run: GSres <- FRBmultiregGS(cbind(reading,mathematics,selfesteem)~., data=schooldata,
bdp = 0.25, conf = 0.8,R=99)
## End(Not run)
#the print method just displays the coefficient estimates
GSres
#the summary function additionally displays the bootstrap standard errors and p-values
#("BCA" method by default)
summary(GSres)
summary(GSres, confmethod="basic")
#ask explicitely for the coefficient matrix:
GSres$coefficients
# or equivalently,
coef(GSres)
#For the error covariance matrix:
GSres$Sigma
#plot some bootstrap histograms for the coefficient estimates
#(with "BCA" intervals by default)
plot(GSres, expl=c("education", "occupation"), resp=c("selfesteem","reading"))
#plot bootstrap histograms for all coefficient estimates
plot(GSres)
#possibly the plot-function has made a selection of coefficients to plot here,
#since 'all' may have been too many to fit on one page, see help(plot.FRBmultireg);
#this is platform-dependent
Results
R version 3.3.1 (2016-06-21) -- "Bug in Your Hair"
Copyright (C) 2016 The R Foundation for Statistical Computing
Platform: x86_64-pc-linux-gnu (64-bit)
R is free software and comes with ABSOLUTELY NO WARRANTY.
You are welcome to redistribute it under certain conditions.
Type 'license()' or 'licence()' for distribution details.
R is a collaborative project with many contributors.
Type 'contributors()' for more information and
'citation()' on how to cite R or R packages in publications.
Type 'demo()' for some demos, 'help()' for on-line help, or
'help.start()' for an HTML browser interface to help.
Type 'q()' to quit R.
> library(FRB)
Loading required package: corpcor
Loading required package: rrcov
Loading required package: robustbase
Scalable Robust Estimators with High Breakdown Point (version 1.3-11)
> png(filename="/home/ddbj/snapshot/RGM3/R_CC/result/FRB/FRBmultiregGS.Rd_%03d_medium.png", width=480, height=480)
> ### Name: FRBmultiregGS
> ### Title: GS-Estimates for multivariate regression with bootstrap
> ### confidence intervals
> ### Aliases: FRBmultiregGS FRBmultiregGS.default FRBmultiregGS.formula
> ### Keywords: multivariate robust
>
> ### ** Examples
>
> data(schooldata)
> school.x <- data.matrix(schooldata[,1:5])
> school.y <- data.matrix(schooldata[,6:8])
>
> #computes 25% breakdown point GS-estimate and 80% confidence intervals
> #based on 99 bootstrap samples:
> GSres <- FRBmultiregGS(school.x, school.y, R=99, bdp = 0.25, conf = 0.8,nsamp=50)
> #or using the formula interface
> ## Not run:
> ##D GSres <- FRBmultiregGS(cbind(reading,mathematics,selfesteem)~., data=schooldata,
> ##D bdp = 0.25, conf = 0.8,R=99)
> ## End(Not run)
>
> #the print method just displays the coefficient estimates
> GSres
Multivariate regression based on multivariate GS-estimates (breakdown point = 0.25)
Coefficients:
reading mathematics selfesteem
(intercept) 1.9406 2.4878 0.0754
education 0.1218 0.0124 -0.0467
occupation 4.9100 6.0603 2.1394
visit 0.0675 0.0197 0.2377
counseling -0.7759 -0.8136 -0.0814
teacher -0.1876 -0.2889 0.0228
>
> #the summary function additionally displays the bootstrap standard errors and p-values
> #("BCA" method by default)
> summary(GSres)
Multivariate regression based on GS-estimates (breakdown point = 0.25)
Response reading:
Residuals:
Min 1Q Median 3Q Max
-14.697 -1.860 0.258 2.359 26.800
Coefficients:
Estimate Std.Error p-value
(intercept) 1.9406 1.2097 0.0904 .
education 0.1218 1.2795 0.0000 ***
occupation 4.9100 0.3818 0.8898
visit 0.0675 0.0727 0.0830 .
counseling -0.7759 1.2053 0.0000 ***
teacher -0.1876 0.3265 0.8331
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
p-values based on BCA method!
Response mathematics:
Residuals:
Min 1Q Median 3Q Max
-10.505 -2.837 -0.573 3.122 38.599
Coefficients:
Estimate Std.Error p-value
(intercept) 2.4878 0.214 0.0000 ***
education 0.0124 0.153 0.1354
occupation 6.0603 0.085 0.9282
visit 0.0197 1.200 0.0000 ***
counseling -0.8136 0.298 0.9596
teacher -0.2889 0.320 0.0451 *
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
p-values based on BCA method!
Response selfesteem:
Residuals:
Min 1Q Median 3Q Max
-2.6744 -0.6642 0.0195 0.6830 3.4961
Coefficients:
Estimate Std.Error p-value
(intercept) 0.0754 0.1946 0.123
education -0.0467 0.0467 0.467
occupation 2.1394 0.6071 0.000 ***
visit 0.2377 0.0920 0.000 ***
counseling -0.0814 0.1330 0.462
teacher 0.0228 0.0460 0.625
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
p-values based on BCA method!
Robust residual scale: 2.22
Error covariance matrix estimate:
reading mathematics selfesteem
reading 14.8 14.59 2.60
mathematics 14.6 21.59 2.65
selfesteem 2.6 2.65 1.59
>
> summary(GSres, confmethod="basic")
Multivariate regression based on GS-estimates (breakdown point = 0.25)
Response reading:
Residuals:
Min 1Q Median 3Q Max
-14.697 -1.860 0.258 2.359 26.800
Coefficients:
Estimate Std.Error p-value
(intercept) 1.9406 1.2097 0.182
education 0.1218 1.2795 0.101
occupation 4.9100 0.3818 0.909
visit 0.0675 0.0727 0.202
counseling -0.7759 1.2053 0.000 ***
teacher -0.1876 0.3265 0.909
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
p-values based on Basic Bootstrap!
Response mathematics:
Residuals:
Min 1Q Median 3Q Max
-10.505 -2.837 -0.573 3.122 38.599
Coefficients:
Estimate Std.Error p-value
(intercept) 2.4878 0.214 0.000 ***
education 0.0124 0.153 0.202
occupation 6.0603 0.085 0.949
visit 0.0197 1.200 0.000 ***
counseling -0.8136 0.298 0.949
teacher -0.2889 0.320 0.000 ***
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
p-values based on Basic Bootstrap!
Response selfesteem:
Residuals:
Min 1Q Median 3Q Max
-2.6744 -0.6642 0.0195 0.6830 3.4961
Coefficients:
Estimate Std.Error p-value
(intercept) 0.0754 0.1946 0.2424
education -0.0467 0.0467 0.0808 .
occupation 2.1394 0.6071 0.0000 ***
visit 0.2377 0.0920 0.0000 ***
counseling -0.0814 0.1330 0.4646
teacher 0.0228 0.0460 0.4444
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
p-values based on Basic Bootstrap!
Robust residual scale: 2.22
Error covariance matrix estimate:
reading mathematics selfesteem
reading 14.8 14.59 2.60
mathematics 14.6 21.59 2.65
selfesteem 2.6 2.65 1.59
>
> #ask explicitely for the coefficient matrix:
> GSres$coefficients
reading mathematics selfesteem
(intercept) 1.94063330 2.48784629 0.07544784
education 0.12180089 0.01239441 -0.04673212
occupation 4.90995552 6.06025859 2.13937389
visit 0.06746034 0.01970355 0.23773282
counseling -0.77593115 -0.81356534 -0.08140681
teacher -0.18758042 -0.28890086 0.02275939
> # or equivalently,
> coef(GSres)
reading mathematics selfesteem
(intercept) 1.94063330 2.48784629 0.07544784
education 0.12180089 0.01239441 -0.04673212
occupation 4.90995552 6.06025859 2.13937389
visit 0.06746034 0.01970355 0.23773282
counseling -0.77593115 -0.81356534 -0.08140681
teacher -0.18758042 -0.28890086 0.02275939
> #For the error covariance matrix:
> GSres$Sigma
reading mathematics selfesteem
reading 14.775899 14.594412 2.598958
mathematics 14.594412 21.590800 2.647048
selfesteem 2.598958 2.647048 1.586931
>
> #plot some bootstrap histograms for the coefficient estimates
> #(with "BCA" intervals by default)
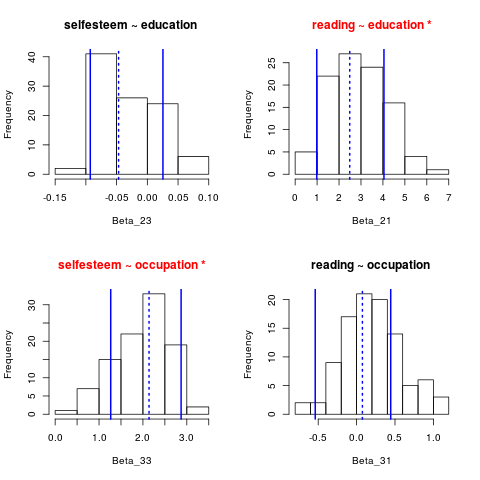
> plot(GSres, expl=c("education", "occupation"), resp=c("selfesteem","reading"))
>
> #plot bootstrap histograms for all coefficient estimates
> plot(GSres)
Warning message:
In plot.FRBmultireg(GSres) :
Number of plots too large to fit on the page, subset was selected: consider specifying (fewer)
variables in 'expl' and 'resp'; or enlarge graphics device; or set 'onepage=FALSE'
> #possibly the plot-function has made a selection of coefficients to plot here,
> #since 'all' may have been too many to fit on one page, see help(plot.FRBmultireg);
> #this is platform-dependent
>
>
>
>
>
>
> dev.off()
null device
1
>
|