Supported by Dr. Osamu Ogasawara and  providing providing  . . |
|
Last data update: 2014.03.03 |
GammaregDescriptionFunction to do Classic Gamma Regression: joint mean and shape modeling UsageGammareg(formula1,formula2,meanlink) Arguments
DetailsThe classic gamma regression allow the joint modelling of mean and shape parameters of a gamma distributed variable, as is proposed in Cepeda (2001), using the Fisher Socring algorithm, with two differentes link for the mean: log and identity, and log link for the shape. Valueobject of class bayesbetareg with:
Author(s)Martha Corrales martha.corrales@usa.edu.co Edilberto Cepeda-Cuervo ecepedac@unal.edu.co References1. Cepeda-Cuervo, E. (2001). Modelagem da variabilidade em modelos lineares generalizados. Unpublished Ph.D. tesis. Instituto de Matem<c3><a1>ticas. Universidade Federal do R<c3><ad>o do Janeiro. //http://www.docentes.unal.edu.co/ecepedac/docs/MODELAGEM20DA20VARIABILIDADE.pdf. http://www.bdigital.unal.edu.co/9394/. 2. McCullagh, P. and Nelder, N.A. (1989). Generalized Linear Models. Second Edition. Chapman and Hall. Examples
#
num.killed <- c(7,59,115,149,178,229,5,43,76,4,57,83,6,57,84)
size.sam <- c(1,2,3,3,3,3,rep(1,9))*100
insecticide <- c(4,5,8,10,15,20,2,5,10,2,5,10,2,5,10)
insecticide.2 <- insecticide^2
synergist <- c(rep(0,6),rep(3.9,3),rep(19.5,3),rep(39,3))
par(mfrow=c(2,2))
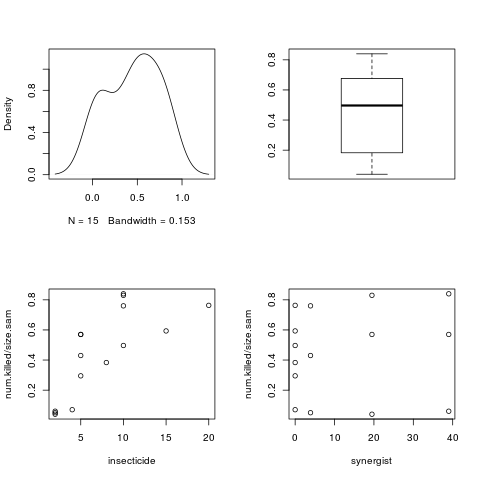
plot(density(num.killed/size.sam),main="")
boxplot(num.killed/size.sam)
plot(insecticide,num.killed/size.sam)
plot(synergist,num.killed/size.sam)
mean.for <- (num.killed/size.sam) ~ insecticide + insecticide.2
dis.for <- ~ synergist + insecticide
res=Gammareg(mean.for,dis.for,meanlink="ide")
summary(glm((num.killed/size.sam) ~ insecticide + insecticide.2,family=Gamma("log")))
summary(res)
# Simulation Example
X1 <- rep(1,500)
X2 <- runif(500,0,30)
X3 <- runif(500,0,15)
X4 <- runif(500,10,20)
mui <- 15 + 2*X2 + 3*X3
alphai <- exp(0.2 + 0.1*X2 + 0.3*X4)
Y <- rgamma(500,shape=alphai,scale=mui/alphai)
X <- cbind(X1,X2,X3)
Z <- cbind(X1,X2,X4)
formula.mean= Y~X2+X3
formula.shape= ~X2+X4
a=Gammareg(formula.mean,formula.shape,meanlink="ide")
summary(a)
Results
R version 3.3.1 (2016-06-21) -- "Bug in Your Hair"
Copyright (C) 2016 The R Foundation for Statistical Computing
Platform: x86_64-pc-linux-gnu (64-bit)
R is free software and comes with ABSOLUTELY NO WARRANTY.
You are welcome to redistribute it under certain conditions.
Type 'license()' or 'licence()' for distribution details.
R is a collaborative project with many contributors.
Type 'contributors()' for more information and
'citation()' on how to cite R or R packages in publications.
Type 'demo()' for some demos, 'help()' for on-line help, or
'help.start()' for an HTML browser interface to help.
Type 'q()' to quit R.
> library(Gammareg)
> png(filename="/home/ddbj/snapshot/RGM3/R_CC/result/Gammareg/Gammareg.Rd_%03d_medium.png", width=480, height=480)
> ### Name: Gammareg
> ### Title: Gammareg
> ### Aliases: Gammareg
> ### Keywords: Classic estimation Fisher Scoring joint modeling Gamma
> ### regression Mean and shape parameters
>
> ### ** Examples
>
>
> #
>
> num.killed <- c(7,59,115,149,178,229,5,43,76,4,57,83,6,57,84)
> size.sam <- c(1,2,3,3,3,3,rep(1,9))*100
> insecticide <- c(4,5,8,10,15,20,2,5,10,2,5,10,2,5,10)
> insecticide.2 <- insecticide^2
> synergist <- c(rep(0,6),rep(3.9,3),rep(19.5,3),rep(39,3))
>
> par(mfrow=c(2,2))
> plot(density(num.killed/size.sam),main="")
> boxplot(num.killed/size.sam)
> plot(insecticide,num.killed/size.sam)
> plot(synergist,num.killed/size.sam)
>
>
> mean.for <- (num.killed/size.sam) ~ insecticide + insecticide.2
> dis.for <- ~ synergist + insecticide
>
> res=Gammareg(mean.for,dis.for,meanlink="ide")
>
> summary(glm((num.killed/size.sam) ~ insecticide + insecticide.2,family=Gamma("log")))
Call:
glm(formula = (num.killed/size.sam) ~ insecticide + insecticide.2,
family = Gamma("log"))
Deviance Residuals:
Min 1Q Median 3Q Max
-0.8439 -0.4926 -0.1526 0.2123 0.8703
Coefficients:
Estimate Std. Error t value Pr(>|t|)
(Intercept) -3.48260 0.47761 -7.292 9.58e-06 ***
insecticide 0.52757 0.11376 4.638 0.000573 ***
insecticide.2 -0.01911 0.00546 -3.500 0.004381 **
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
(Dispersion parameter for Gamma family taken to be 0.3815866)
Null deviance: 11.9764 on 14 degrees of freedom
Residual deviance: 4.1655 on 12 degrees of freedom
AIC: -3.8819
Number of Fisher Scoring iterations: 8
> summary(res)
################################################################
### Classic Gamma Regression ###
################################################################
Call:
Gammareg(formula1 = mean.for, formula2 = dis.for, meanlink = "ide")
Estimate L.Intv U.Intv
beta.(Intercept) -0.129697 -0.261399 0.002
beta.insecticide 0.119233 0.068680 0.170
beta.insecticide.2 -0.004089 -0.006540 -0.002
gamma.(Intercept) 0.439037 0.182929 0.695
gamma.synergist 0.015735 0.010504 0.021
gamma.insecticide 0.162918 0.149205 0.177
Covariance Matrix for Beta:
(Intercept) insecticide insecticide.2
(Intercept) 3.580541e-03 -1.213089e-03 5.459089e-05
insecticide -1.213089e-03 5.275563e-04 -2.494403e-05
insecticide.2 5.459089e-05 -2.494403e-05 1.239960e-06
Covariance Matrix for Gamma:
(Intercept) synergist insecticide
(Intercept) 0.0138168350 -2.122943e-04 -7.266857e-04
synergist -0.0002122943 5.764239e-06 1.080063e-05
insecticide -0.0007266857 1.080063e-05 3.961207e-05
AIC:
[1] -577.8824
Iteration:
[1] 59
Convergence:
[1] 5.61794e-06
>
> # Simulation Example
>
> X1 <- rep(1,500)
> X2 <- runif(500,0,30)
> X3 <- runif(500,0,15)
> X4 <- runif(500,10,20)
> mui <- 15 + 2*X2 + 3*X3
> alphai <- exp(0.2 + 0.1*X2 + 0.3*X4)
> Y <- rgamma(500,shape=alphai,scale=mui/alphai)
> X <- cbind(X1,X2,X3)
> Z <- cbind(X1,X2,X4)
> formula.mean= Y~X2+X3
> formula.shape= ~X2+X4
> a=Gammareg(formula.mean,formula.shape,meanlink="ide")
> summary(a)
################################################################
### Classic Gamma Regression ###
################################################################
Call:
Gammareg(formula1 = formula.mean, formula2 = formula.shape, meanlink = "ide")
Estimate L.Intv U.Intv
beta.(Intercept) 15.3913 14.9044 15.878
beta.X2 2.0001 1.9763 2.024
beta.X3 2.9851 2.9399 3.030
gamma.(Intercept) 0.3195 0.3188 0.320
gamma.X2 0.1048 0.1048 0.105
gamma.X4 0.2866 0.2866 0.287
Covariance Matrix for Beta:
(Intercept) X2 X3
(Intercept) 0.061409236 -2.264497e-03 -2.365192e-03
X2 -0.002264497 1.460311e-04 -3.309234e-05
X3 -0.002365192 -3.309234e-05 5.280078e-04
Covariance Matrix for Gamma:
(Intercept) X2 X4
(Intercept) 1.311470e-07 -1.008708e-09 -5.763330e-09
X2 -1.008708e-09 3.772879e-11 4.617491e-12
X4 -5.763330e-09 4.617491e-12 3.084264e-10
AIC:
[1] 7113416
Iteration:
[1] 11
Convergence:
[1] 3.365878e-10
>
>
>
>
>
>
> dev.off()
null device
1
>
|