Supported by Dr. Osamu Ogasawara and  providing providing  . . |
|
Last data update: 2014.03.03 |
Microarray Time Series Data for 1444 Caulobacter Crescentus GenesDescriptionThis data set describes the temporal expression of 1444 genes (open reading frames) in the cell cycle of the bacterium Caulobacter crescentus. Usagedata(caulobacter) Format
SourceThis data is described in Laub et al. (2000) and can be freely downloaded from (http://caulobacter.stanford.edu/CellCycle/DownloadData.htm). ReferencesLaub, M.T., McAdams, H.H., Feldblyum, Fraser, C.M., and Shapiro, L. (2000) Global analysis of the genetic network controlling a bacterial cell cycle. Science, 290, 2144–1248. Examples
# load GeneCycle library
library("GeneCycle")
# load data set
data(caulobacter)
is.longitudinal(caulobacter)
# how many samples and how many genes?
dim(caulobacter)
summary(caulobacter)
get.time.repeats(caulobacter)
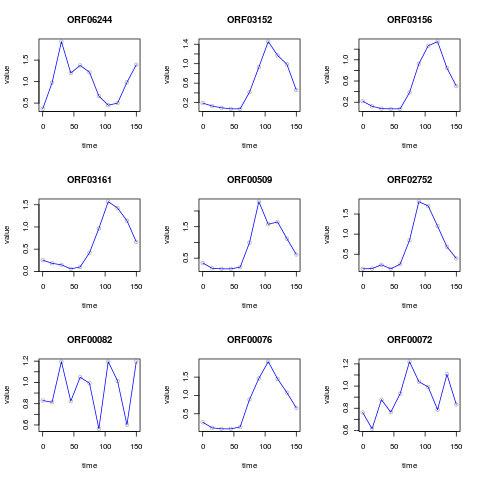
# plot first nine time series
plot(caulobacter, 1:9)
Results
R version 3.3.1 (2016-06-21) -- "Bug in Your Hair"
Copyright (C) 2016 The R Foundation for Statistical Computing
Platform: x86_64-pc-linux-gnu (64-bit)
R is free software and comes with ABSOLUTELY NO WARRANTY.
You are welcome to redistribute it under certain conditions.
Type 'license()' or 'licence()' for distribution details.
R is a collaborative project with many contributors.
Type 'contributors()' for more information and
'citation()' on how to cite R or R packages in publications.
Type 'demo()' for some demos, 'help()' for on-line help, or
'help.start()' for an HTML browser interface to help.
Type 'q()' to quit R.
> library(GeneCycle)
Loading required package: MASS
Loading required package: longitudinal
Loading required package: corpcor
Loading required package: fdrtool
> png(filename="/home/ddbj/snapshot/RGM3/R_CC/result/GeneCycle/caulobacter.Rd_%03d_medium.png", width=480, height=480)
> ### Name: caulobacter
> ### Title: Microarray Time Series Data for 1444 Caulobacter Crescentus
> ### Genes
> ### Aliases: caulobacter
> ### Keywords: datasets
>
> ### ** Examples
>
> # load GeneCycle library
> library("GeneCycle")
>
> # load data set
> data(caulobacter)
> is.longitudinal(caulobacter)
[1] TRUE
>
> # how many samples and how many genes?
> dim(caulobacter)
[1] 11 1444
> summary(caulobacter)
Longitudinal data:
1444 variables measured at 11 different time points
Total number of measurements per variable: 11
Repeated measurements: none
To obtain the measurement design call 'get.time.repeats()'.
> get.time.repeats(caulobacter)
$time
[1] 0 15 30 45 60 75 90 105 120 135 150
$repeats
[1] 1 1 1 1 1 1 1 1 1 1 1
>
> # plot first nine time series
> plot(caulobacter, 1:9)
>
>
>
>
>
> dev.off()
null device
1
>
|