Supported by Dr. Osamu Ogasawara and  providing providing  . . |
|
Last data update: 2014.03.03 |
Mixture Discriminant Analysis with HD GaussiansDescriptionHD-MDA implements mixture discriminant analysis (MDA, Hastie & Tibshirani, 1996) with HD Gaussians instead of full Gaussians. Each class is assumed to be made of several class-specific groups in which the data live in low-dimensional subspaces. From a technical point of view, a clustering is done using Usagehdmda(X,cls,K=1:10,model='AkjBkQkDk',show=FALSE,...) Arguments
DetailsSome information on the signification of the model names:
The model “all” will compute all the models, give their BIC and keep the model with the highest BIC value. Instead of writing the model names, they can also be specified using an integer. 1 represents the most general model (“AkjBkQkDk”) while 14 is the most constrained (“ABQD”), the others number/name matching are given below:
Value
Author(s)Laurent Berge, Charles Bouveyron and Stephane Girard ReferencesC. Bouveyron and C. Brunet (2014), “Model-based clustering of high-dimensional data: A review”, Computational Statistics and Data Analysis, vol. 71, pp. 52-78. Bouveyron, C. Girard, S. and Schmid, C. (2007), “High Dimensional Discriminant Analysis”, Communications in Statistics: Theory and Methods, vol. 36 (14), pp. 2607-2623. Bouveyron, C. Celeux, G. and Girard, S. (2011), “Intrinsic dimension estimation by maximum likelihood in probabilistic PCA”, Pattern Recognition Letters, vol. 32 (14), pp. 1706-1713. Berge, L. Bouveyron, C. and Girard, S. (2012), “HDclassif: An R Package for Model-Based Clustering and Discriminant Analysis of High-Dimensional Data”, Journal of Statistical Software, 46(6), pp. 1-29, url: http://www.jstatsoft.org/v46/i06/. Hastie, T., & Tibshirani, R. (1996), “Discriminant analysis by Gaussian mixtures”, Journal of the Royal Statistical Society, Series B (Methodological), pp. 155-176. See Also
Examples
# Load the Wine data set
data(wine)
cls = wine[,1]; X = scale(wine[,-1])
# A simple use...
out = hdmda(X[1:100,],cls[1:100])
res = predict(out,X[101:nrow(X),])
# Comparison between hdmda and hdda in a CV setup
set.seed(123); nb = 10; Err = matrix(NA,2,nb)
for (i in 1:nb){
cat('.')
test = sample(nrow(X),50)
out0 = lda(X[-test,],cls[-test])
res0 = predict(out0,X[test,])
Err[1,i] = sum(res0$class != cls[test]) / length(test)
out = hdmda(X[-test,],cls[-test],K=1:3,model="AKJBQKDK")
res = predict(out,X[test,])
Err[2,i] = sum(res$class != cls[test]) / length(test)
}
cat('\n')
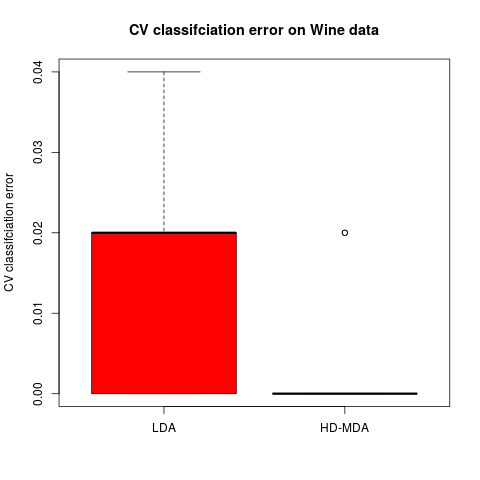
boxplot(t(Err),names=c('LDA','HD-MDA'),col=2:3,ylab="CV classifciation error",
main='CV classifciation error on Wine data')
Results
R version 3.3.1 (2016-06-21) -- "Bug in Your Hair"
Copyright (C) 2016 The R Foundation for Statistical Computing
Platform: x86_64-pc-linux-gnu (64-bit)
R is free software and comes with ABSOLUTELY NO WARRANTY.
You are welcome to redistribute it under certain conditions.
Type 'license()' or 'licence()' for distribution details.
R is a collaborative project with many contributors.
Type 'contributors()' for more information and
'citation()' on how to cite R or R packages in publications.
Type 'demo()' for some demos, 'help()' for on-line help, or
'help.start()' for an HTML browser interface to help.
Type 'q()' to quit R.
> library(HDclassif)
Loading required package: MASS
> png(filename="/home/ddbj/snapshot/RGM3/R_CC/result/HDclassif/hdmda.Rd_%03d_medium.png", width=480, height=480)
> ### Name: hdmda
> ### Title: Mixture Discriminant Analysis with HD Gaussians
> ### Aliases: hdmda
> ### Keywords: mixture discriminant analysis high-dimensional data
>
> ### ** Examples
>
> # Load the Wine data set
> data(wine)
> cls = wine[,1]; X = scale(wine[,-1])
>
> # A simple use...
> out = hdmda(X[1:100,],cls[1:100])
> res = predict(out,X[101:nrow(X),])
>
> # Comparison between hdmda and hdda in a CV setup
> set.seed(123); nb = 10; Err = matrix(NA,2,nb)
> for (i in 1:nb){
+ cat('.')
+ test = sample(nrow(X),50)
+ out0 = lda(X[-test,],cls[-test])
+ res0 = predict(out0,X[test,])
+ Err[1,i] = sum(res0$class != cls[test]) / length(test)
+ out = hdmda(X[-test,],cls[-test],K=1:3,model="AKJBQKDK")
+ res = predict(out,X[test,])
+ Err[2,i] = sum(res$class != cls[test]) / length(test)
+ }
..........> cat('\n')
> boxplot(t(Err),names=c('LDA','HD-MDA'),col=2:3,ylab="CV classifciation error",
+ main='CV classifciation error on Wine data')
>
>
>
>
>
> dev.off()
null device
1
>
|