Supported by Dr. Osamu Ogasawara and  providing providing  . . |
|
Last data update: 2014.03.03 |
Functions for transdimensional MCMCDescriptionComputes the value of the 'g' density at a given point and, optionally, returns the backtransformed point and the model to which the point belongs. Usagetrans.dens(y, ldens.list, which.models, ..., back.transform=F) trans.up(x, ldens.list, which.models, ...) trans2(y, ldens.list, k, ...) transUp2(y, ldens.list, k, ...) transBack2(y, ldens.list, k, ...) Arguments
DetailsSee the reference for details. The functions with the 2 in the name operate on pairs of models only. ValueIf Author(s)Giovanni Petris GPetris@uark.edu, Luca Tardella ReferencesPetris & Tardella, A geometric approach to transdimensional Markov chain Monte Carlo. The Canadian Journal of Statistics, vol.31, n.4, (2003). Examples
#### ==> Warning: running the examples may take a few minutes! <== ####
### Generate a sample from a mixture of 0,1,2-dim standard normals
ldens.list <- list(f0 = function(x) sum(dnorm(x,log=TRUE)),
f1 = function(x) dnorm(x,log=TRUE),
f2 = function() 0)
trans.mix <- function(y) {
trans.dens(y, ldens.list=ldens.list, which.models=0:2)
}
trans.rmix <- arms(c(0,0), trans.mix, function(x) crossprod(x)<1e4, 500)
rmix <- trans.dens(y=trans.rmix, ldens.list=ldens.list,
which.models=0:2, back.transform = TRUE)
table(rmix[,2])/nrow(rmix) # should be about equally distributed
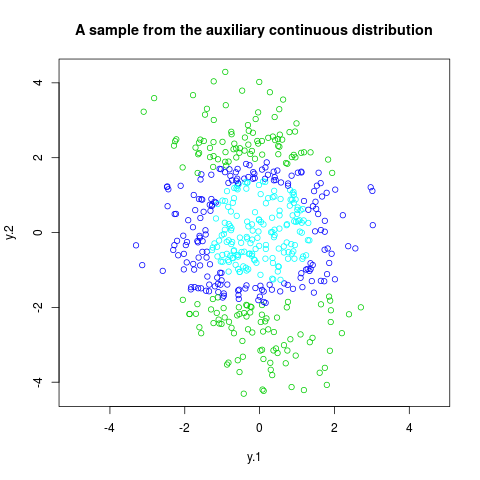
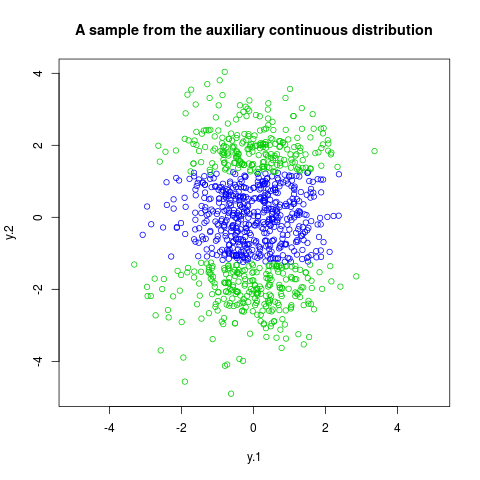
plot(trans.rmix,col=rmix[,2]+3,asp=1, xlab="y.1", ylab="y.2",
main="A sample from the auxiliary continuous distribution")
x <- rmix[,-(1:2)]
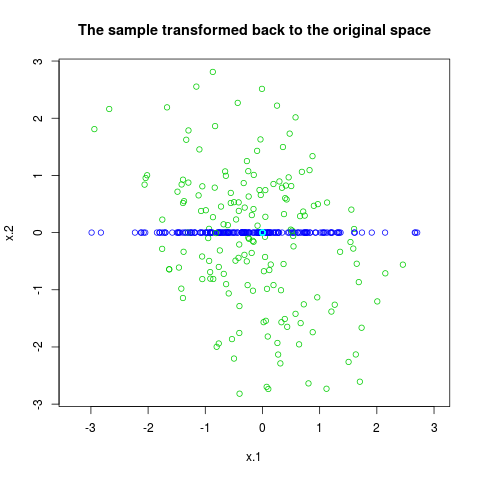
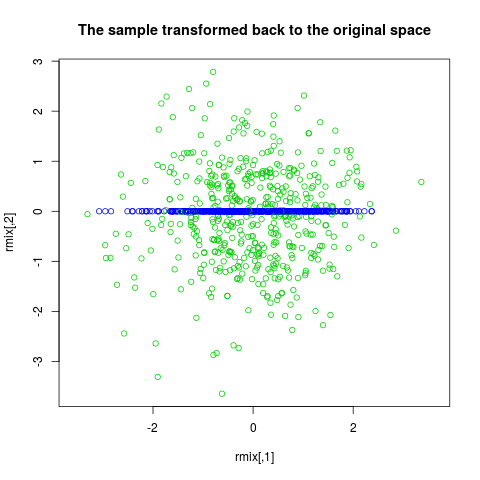
plot(x, col=rmix[,2]+3, asp=1,
main="The sample transformed back to the original space")
### trans.up as a right inverse of trans.dens
set.seed(6324)
y <- trans.up(x, ldens.list, 0:2)
stopifnot(all.equal(x, trans.dens(y, ldens.list, 0:2, back.transform=TRUE)[,-(1:2)]))
### More trans.up
z <- trans.up(matrix(0,1000,2), ldens.list, 0:2)
plot(z,asp=1,col=5) # should look uniform in a circle corresponding to model 2
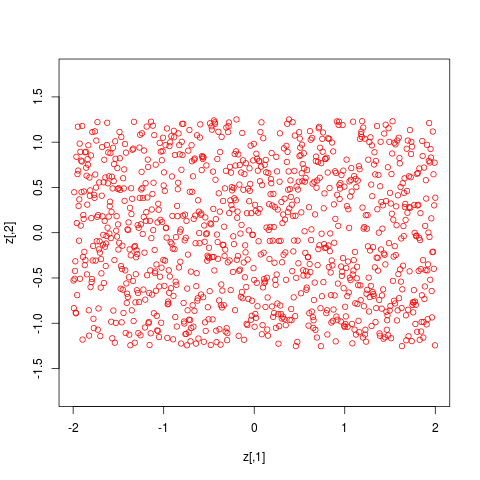
z <- trans.up(cbind(runif(1000,-3,3),0), ldens.list, 0:2)
plot(z,asp=1,col=4) # should look uniform in a region corresponding to model 1
### trans2, transBack2
ldens.list <- list(f0 = function(x) sum(dnorm(x,log=TRUE)),
f1 = function(x) dnorm(x,log=TRUE))
trans.mix <- function(y) {
trans2(y, ldens.list=ldens.list, k=1)[-2]
}
trans.rmix <- arms(c(0,0), trans.mix, function(x) crossprod(x)<1e2, 1000)
rmix <- transBack2(y=trans.rmix, ldens.list=ldens.list, k=1)
table(rmix[,2]==0)/nrow(rmix) # should be about equally distributed
plot(trans.rmix,col=(rmix[,2]==0)+3,asp=1, xlab="y.1", ylab="y.2",
main="A sample from the auxiliary continuous distribution")
plot(rmix, col=(rmix[,2]==0)+3, asp=1,
main="The sample transformed back to the original space")
### trunsUp2
z <- t(sapply(1:1000, function(i) transUp2(c(-2+0.004*i,0), ldens.list, 1)))
plot(z,asp=1,col=2)
Results
R version 3.3.1 (2016-06-21) -- "Bug in Your Hair"
Copyright (C) 2016 The R Foundation for Statistical Computing
Platform: x86_64-pc-linux-gnu (64-bit)
R is free software and comes with ABSOLUTELY NO WARRANTY.
You are welcome to redistribute it under certain conditions.
Type 'license()' or 'licence()' for distribution details.
R is a collaborative project with many contributors.
Type 'contributors()' for more information and
'citation()' on how to cite R or R packages in publications.
Type 'demo()' for some demos, 'help()' for on-line help, or
'help.start()' for an HTML browser interface to help.
Type 'q()' to quit R.
> library(HI)
> png(filename="/home/ddbj/snapshot/RGM3/R_CC/result/HI/trans.dens.Rd_%03d_medium.png", width=480, height=480)
> ### Name: trans.dens
> ### Title: Functions for transdimensional MCMC
> ### Aliases: trans.dens trans.up trans2 transUp2 transBack2
> ### Keywords: distribution
>
> ### ** Examples
>
> #### ==> Warning: running the examples may take a few minutes! <== ####
> ### Generate a sample from a mixture of 0,1,2-dim standard normals
> ldens.list <- list(f0 = function(x) sum(dnorm(x,log=TRUE)),
+ f1 = function(x) dnorm(x,log=TRUE),
+ f2 = function() 0)
> trans.mix <- function(y) {
+ trans.dens(y, ldens.list=ldens.list, which.models=0:2)
+ }
>
> trans.rmix <- arms(c(0,0), trans.mix, function(x) crossprod(x)<1e4, 500)
> rmix <- trans.dens(y=trans.rmix, ldens.list=ldens.list,
+ which.models=0:2, back.transform = TRUE)
> table(rmix[,2])/nrow(rmix) # should be about equally distributed
0 1 2
0.330 0.348 0.322
> plot(trans.rmix,col=rmix[,2]+3,asp=1, xlab="y.1", ylab="y.2",
+ main="A sample from the auxiliary continuous distribution")
> x <- rmix[,-(1:2)]
> plot(x, col=rmix[,2]+3, asp=1,
+ main="The sample transformed back to the original space")
> ### trans.up as a right inverse of trans.dens
> set.seed(6324)
> y <- trans.up(x, ldens.list, 0:2)
> stopifnot(all.equal(x, trans.dens(y, ldens.list, 0:2, back.transform=TRUE)[,-(1:2)]))
>
> ### More trans.up
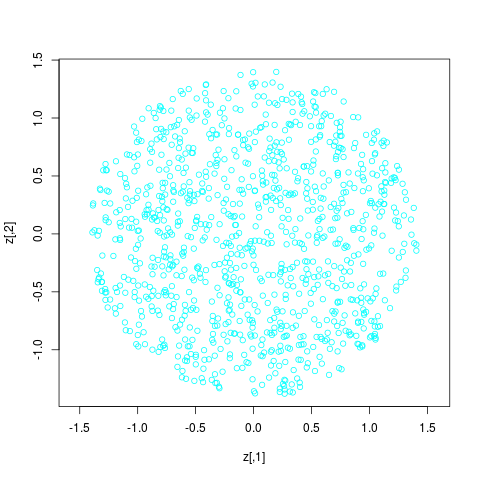
> z <- trans.up(matrix(0,1000,2), ldens.list, 0:2)
> plot(z,asp=1,col=5) # should look uniform in a circle corresponding to model 2
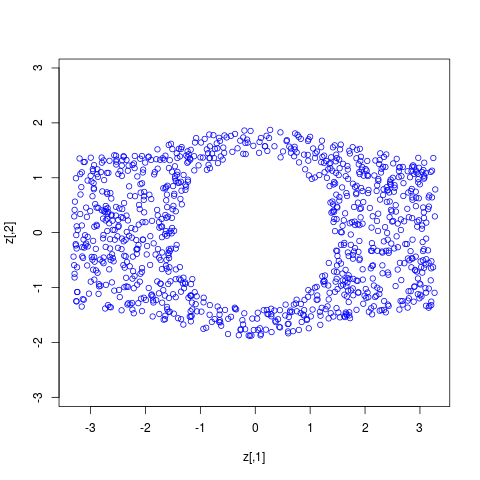
> z <- trans.up(cbind(runif(1000,-3,3),0), ldens.list, 0:2)
> plot(z,asp=1,col=4) # should look uniform in a region corresponding to model 1
>
> ### trans2, transBack2
> ldens.list <- list(f0 = function(x) sum(dnorm(x,log=TRUE)),
+ f1 = function(x) dnorm(x,log=TRUE))
> trans.mix <- function(y) {
+ trans2(y, ldens.list=ldens.list, k=1)[-2]
+ }
> trans.rmix <- arms(c(0,0), trans.mix, function(x) crossprod(x)<1e2, 1000)
> rmix <- transBack2(y=trans.rmix, ldens.list=ldens.list, k=1)
> table(rmix[,2]==0)/nrow(rmix) # should be about equally distributed
FALSE TRUE
0.496 0.504
> plot(trans.rmix,col=(rmix[,2]==0)+3,asp=1, xlab="y.1", ylab="y.2",
+ main="A sample from the auxiliary continuous distribution")
> plot(rmix, col=(rmix[,2]==0)+3, asp=1,
+ main="The sample transformed back to the original space")
>
> ### trunsUp2
> z <- t(sapply(1:1000, function(i) transUp2(c(-2+0.004*i,0), ldens.list, 1)))
> plot(z,asp=1,col=2)
>
>
>
>
>
>
> dev.off()
null device
1
>
|