Supported by Dr. Osamu Ogasawara and  providing providing  . . |
|
Last data update: 2014.03.03 |
Lilliefors-corrected Kolmogorov-Smirnoff Goodness-Of-Fit TestDescriptionImplements the Lilliefors-corrected Kolmogorov-Smirnoff test for use in
goodness-of-fit tests, suitable when population parameters are unknown and
must be estimated by sample statistics. It uses Monte Carlo simulation to
estimate p-values. Coded to wrap around UsageLcKS(x, cdf, nreps = 4999, G = 1:9) Arguments
DetailsThe function builds a simulation distribution The default The p-value is calculated as the number of Monte Carlo samples with
test statistics D as extreme as or more extreme than that in the
observed sample Parameter estimates are calculated for the specified continuous
distribution, using maximum-likelihood estimates. When testing against the
gamma and Weibull distributions, Sample statistics for the (univariate) normal mixture distribution
Be aware that constraining The ValueA list containing the following components:
NoteThe Kolmogorov-Smirnoff (such as Author(s)Phil Novack-Gottshall pnovack-gottshall@ben.edu, based on code from Charles Geyer (University of Minnesota). ReferencesGihman, I. I. 1952. Ob empiriceskoj funkcii raspredelenija slucaje grouppirovki dannych [On the empirical distribution function in the case of grouping of observations]. Doklady Akademii Nauk SSSR 82: 837-840. Lilliefors, H. W. 1967. On the Kolmogorov-Smirnov test for normality with mean and variance unknown. Journal of the American Statistical Association 62(318):399-402. Lilliefors, H. W. 1969. On the Kolmogorov-Smirnov test for the exponential distribution with mean unknown. Journal of the American Statistical Association 64(325):387-389. Manly, B. F. J. 2004. Randomization, Bootstrap and Monte Carlo Methods in Biology. Chapman & Hall, Cornwall, Great Britain. Parsons, F. G., and P. H. Wirsching. 1982. A Kolmogorov-Smirnov goodness-of-fit test for the two-parameter Weibull distribution when the parameters are estimated from the data. Microelectronics Reliability 22(2):163-167. See Also
Examplesx <- runif(200) Lc <- LcKS(x, cdf="pnorm", nreps=999) hist(Lc$D.sim) abline(v = Lc$D.obs, lty = 2) print(Lc, max=50) # Print first 50 simulated statistics # Approximate p-value (usually) << 0.05 # Confirmation uncorrected version has increased Type II error rate when # using sample statistics to estimate parameters: ks.test(x, "pnorm", mean(x), sd(x)) # p-value always larger, (usually) > 0.05 # Confirm critical values for normal distribution are correct nreps <- 9999 x <- rnorm(25) Lc <- LcKS(x, "pnorm", nreps=nreps) sim.Ds <- sort(Lc$D.sim) crit <- round(c(.8, .85, .9, .95, .99) * nreps, 0) # Lilliefors' (1967) critical values, using improved values from # Parsons & Wirsching (1982) (for n=25): # 0.141 0.148 0.157 0.172 0.201 round(sim.Ds[crit], 3) # Approximately the same critical values # Confirm critical values for exponential are the same as reported by Lilliefors (1969) nreps <- 9999 x <- rexp(25) Lc <- LcKS(x, "pexp", nreps=nreps) sim.Ds <- sort(Lc$D.sim) crit <- round(c(.8, .85, .9, .95, .99) * nreps, 0) # Lilliefors' (1969) critical values (for n=25): # 0.170 0.180 0.191 0.210 0.247 round(sim.Ds[crit], 3) # Approximately the same critical values ## Not run: # Gamma and Weibull tests require functions from the 'MASS' package # Takes time for maximum likelihood optimization of statistics require(MASS) x <- runif(100, min=1, max=100) Lc <- LcKS(x, cdf="pgamma", nreps=499) Lc$p.value # Confirm critical values for Weibull the same as reported by Parsons & Wirsching (1982) nreps <- 9999 x <- rweibull(25, shape=1, scale=1) Lc <- LcKS(x, "pweibull", nreps=nreps) sim.Ds <- sort(Lc$D.sim) crit <- round(c(.8, .85, .9, .95, .99) * nreps, 0) # Parsons & Wirsching (1982) critical values (for n=25): # 0.141 0.148 0.157 0.172 0.201 round(sim.Ds[crit], 3) # Approximately the same critical values # Mixture test requires functions from the 'mclust' package # Takes time to identify model parameters require(mclust) x <- rmixnorm(200, mean=c(10, 20), sd=2, pro=c(1,3)) Lc <- LcKS(x, cdf="pmixnorm", nreps=499, G=1:9) # Default G (1:9) takes long time Lc$p.value G <- Mclust(x)$parameters$variance$G # Optimal model has only two components Lc <- LcKS(x, cdf="pmixnorm", nreps=499, G=G) # Restricting to likely G saves time # But note changes null hypothesis: now testing against just two-component mixture Lc$p.value ## End(Not run) Results
R version 3.3.1 (2016-06-21) -- "Bug in Your Hair"
Copyright (C) 2016 The R Foundation for Statistical Computing
Platform: x86_64-pc-linux-gnu (64-bit)
R is free software and comes with ABSOLUTELY NO WARRANTY.
You are welcome to redistribute it under certain conditions.
Type 'license()' or 'licence()' for distribution details.
R is a collaborative project with many contributors.
Type 'contributors()' for more information and
'citation()' on how to cite R or R packages in publications.
Type 'demo()' for some demos, 'help()' for on-line help, or
'help.start()' for an HTML browser interface to help.
Type 'q()' to quit R.
> library(KScorrect)
> png(filename="/home/ddbj/snapshot/RGM3/R_CC/result/KScorrect/LcKS.Rd_%03d_medium.png", width=480, height=480)
> ### Name: LcKS
> ### Title: Lilliefors-corrected Kolmogorov-Smirnoff Goodness-Of-Fit Test
> ### Aliases: LcKS
>
> ### ** Examples
>
> x <- runif(200)
> Lc <- LcKS(x, cdf="pnorm", nreps=999)
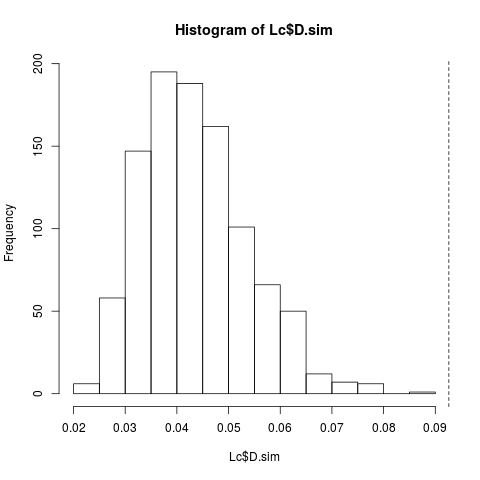
> hist(Lc$D.sim)
> abline(v = Lc$D.obs, lty = 2)
> print(Lc, max=50) # Print first 50 simulated statistics
$D.obs
[1] 0.09262385
$D.sim
[1] 0.03706487 0.03842167 0.02583074 0.04725934 0.04539004 0.02914926
[7] 0.04583008 0.04229889 0.04126987 0.03473239 0.03666469 0.05564759
[13] 0.04883253 0.03809087 0.03073768 0.05027152 0.04581409 0.03636533
[19] 0.03939353 0.02878655 0.05871600 0.03299456 0.07644536 0.05299444
[25] 0.02932849 0.04356083 0.05483117 0.03369578 0.03039681 0.06930097
[31] 0.04429688 0.03832761 0.06029079 0.06807311 0.04295805 0.06358311
[37] 0.05347978 0.03548766 0.02621571 0.03770416 0.03833268 0.04038596
[43] 0.06787367 0.03784615 0.03743014 0.04957043 0.04784248 0.03803586
[49] 0.05131795 0.05864168
[ reached getOption("max.print") -- omitted 949 entries ]
$p.value
[1] 0.001
> # Approximate p-value (usually) << 0.05
>
> # Confirmation uncorrected version has increased Type II error rate when
> # using sample statistics to estimate parameters:
> ks.test(x, "pnorm", mean(x), sd(x)) # p-value always larger, (usually) > 0.05
One-sample Kolmogorov-Smirnov test
data: x
D = 0.092624, p-value = 0.06466
alternative hypothesis: two-sided
>
> # Confirm critical values for normal distribution are correct
> nreps <- 9999
> x <- rnorm(25)
> Lc <- LcKS(x, "pnorm", nreps=nreps)
> sim.Ds <- sort(Lc$D.sim)
> crit <- round(c(.8, .85, .9, .95, .99) * nreps, 0)
> # Lilliefors' (1967) critical values, using improved values from
> # Parsons & Wirsching (1982) (for n=25):
> # 0.141 0.148 0.157 0.172 0.201
> round(sim.Ds[crit], 3) # Approximately the same critical values
[1] 0.142 0.150 0.159 0.173 0.203
>
> # Confirm critical values for exponential are the same as reported by Lilliefors (1969)
> nreps <- 9999
> x <- rexp(25)
> Lc <- LcKS(x, "pexp", nreps=nreps)
> sim.Ds <- sort(Lc$D.sim)
> crit <- round(c(.8, .85, .9, .95, .99) * nreps, 0)
> # Lilliefors' (1969) critical values (for n=25):
> # 0.170 0.180 0.191 0.210 0.247
> round(sim.Ds[crit], 3) # Approximately the same critical values
[1] 0.170 0.179 0.191 0.211 0.251
>
> ## Not run:
> ##D # Gamma and Weibull tests require functions from the 'MASS' package
> ##D # Takes time for maximum likelihood optimization of statistics
> ##D require(MASS)
> ##D x <- runif(100, min=1, max=100)
> ##D Lc <- LcKS(x, cdf="pgamma", nreps=499)
> ##D Lc$p.value
> ##D
> ##D # Confirm critical values for Weibull the same as reported by Parsons & Wirsching (1982)
> ##D nreps <- 9999
> ##D x <- rweibull(25, shape=1, scale=1)
> ##D Lc <- LcKS(x, "pweibull", nreps=nreps)
> ##D sim.Ds <- sort(Lc$D.sim)
> ##D crit <- round(c(.8, .85, .9, .95, .99) * nreps, 0)
> ##D # Parsons & Wirsching (1982) critical values (for n=25):
> ##D # 0.141 0.148 0.157 0.172 0.201
> ##D round(sim.Ds[crit], 3) # Approximately the same critical values
> ##D
> ##D # Mixture test requires functions from the 'mclust' package
> ##D # Takes time to identify model parameters
> ##D require(mclust)
> ##D x <- rmixnorm(200, mean=c(10, 20), sd=2, pro=c(1,3))
> ##D Lc <- LcKS(x, cdf="pmixnorm", nreps=499, G=1:9) # Default G (1:9) takes long time
> ##D Lc$p.value
> ##D G <- Mclust(x)$parameters$variance$G # Optimal model has only two components
> ##D Lc <- LcKS(x, cdf="pmixnorm", nreps=499, G=G) # Restricting to likely G saves time
> ##D # But note changes null hypothesis: now testing against just two-component mixture
> ##D Lc$p.value
> ## End(Not run)
>
>
>
>
>
>
> dev.off()
null device
1
>
|