Supported by Dr. Osamu Ogasawara and  providing providing  . . |
|
Last data update: 2014.03.03 |
Useful data to quickly update kriging mean and varianceDescriptionThis function is used in combination with UsageprecomputeUpdateData(model, integration.points) Arguments
ValueA list with components:
Author(s)Clement Chevalier (IMSV, Switzerland, and IRSN, France) ReferencesChevalier C., Bect J., Ginsbourger D., Vazquez E., Picheny V., Richet Y. (2011), Fast parallel kriging-based stepwise uncertainty reduction with application to the identification of an excursion set ,http://hal.archives-ouvertes.fr/hal-00641108/ Chevalier C., Ginsbourger D. (2012), Corrected Kriging update formulae for batch-sequential data assimilation ,http://arxiv.org/pdf/1203.6452.pdf See Also
Examples
#precomputeUpdateData
set.seed(8)
N <- 9 #number of observations
testfun <- branin
#a 9 points initial design
design <- data.frame( matrix(runif(2*N),ncol=2) )
response <- testfun(design)
#km object with matern3_2 covariance
#params estimated by ML from the observations
model <- km(formula=~., design = design,
response = response,covtype="matern3_2")
#the points where we want to compute prediction (if a point new.x is added to the doe)
n.grid <- 20 #you can run it with 100
x.grid <- y.grid <- seq(0,1,length=n.grid)
newdata <- expand.grid(x.grid,y.grid)
precalc.data <- precomputeUpdateData(model=model,integration.points=newdata)
#now we can compute very quickly kriging covariances
#between these data and any other points
other.x <- matrix(c(0.6,0.6),ncol=2)
pred <- predict_nobias_km(object=model,newdata=other.x,type="UK",se.compute=TRUE)
kn <- computeQuickKrigcov(model=model,integration.points=newdata,X.new=other.x,
precalc.data=precalc.data,F.newdata=pred$F.newdata,
c.newdata=pred$c)
z.grid <- matrix(kn, n.grid, n.grid)
#plots: contour of the criterion, doe points and new point
image(x=x.grid,y=y.grid,z=z.grid,col=grey.colors(10))
contour(x=x.grid,y=y.grid,z=z.grid,15,add=TRUE)
contour(x=x.grid,y=y.grid,z=z.grid,levels=0,add=TRUE,col="blue",lwd=5)
points(design, col="black", pch=17, lwd=4,cex=2)
points(other.x, col="red", pch=17, lwd=4,cex=3)
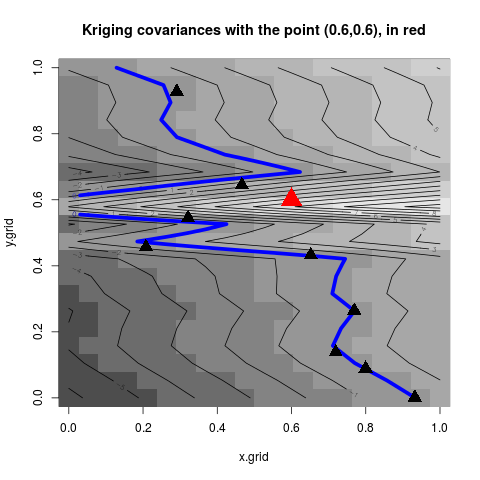
title("Kriging covariances with the point (0.6,0.6), in red")
Results
R version 3.3.1 (2016-06-21) -- "Bug in Your Hair"
Copyright (C) 2016 The R Foundation for Statistical Computing
Platform: x86_64-pc-linux-gnu (64-bit)
R is free software and comes with ABSOLUTELY NO WARRANTY.
You are welcome to redistribute it under certain conditions.
Type 'license()' or 'licence()' for distribution details.
R is a collaborative project with many contributors.
Type 'contributors()' for more information and
'citation()' on how to cite R or R packages in publications.
Type 'demo()' for some demos, 'help()' for on-line help, or
'help.start()' for an HTML browser interface to help.
Type 'q()' to quit R.
> library(KrigInv)
Loading required package: DiceKriging
Loading required package: pbivnorm
Loading required package: rgenoud
## rgenoud (Version 5.7-12.4, Build Date: 2015-07-19)
## See http://sekhon.berkeley.edu/rgenoud for additional documentation.
## Please cite software as:
## Walter Mebane, Jr. and Jasjeet S. Sekhon. 2011.
## ``Genetic Optimization Using Derivatives: The rgenoud package for R.''
## Journal of Statistical Software, 42(11): 1-26.
##
Loading required package: randtoolbox
Loading required package: rngWELL
This is randtoolbox. For overview, type 'help("randtoolbox")'.
> png(filename="/home/ddbj/snapshot/RGM3/R_CC/result/KrigInv/precomputeUpdateData.Rd_%03d_medium.png", width=480, height=480)
> ### Name: precomputeUpdateData
> ### Title: Useful data to quickly update kriging mean and variance
> ### Aliases: precomputeUpdateData
>
> ### ** Examples
>
> #precomputeUpdateData
>
> set.seed(8)
> N <- 9 #number of observations
> testfun <- branin
>
> #a 9 points initial design
> design <- data.frame( matrix(runif(2*N),ncol=2) )
> response <- testfun(design)
>
> #km object with matern3_2 covariance
> #params estimated by ML from the observations
> model <- km(formula=~., design = design,
+ response = response,covtype="matern3_2")
optimisation start
------------------
* estimation method : MLE
* optimisation method : BFGS
* analytical gradient : used
* trend model : ~X1 + X2
* covariance model :
- type : matern3_2
- nugget : NO
- parameters lower bounds : 1e-10 1e-10
- parameters upper bounds : 1.448893 1.853021
- best initial criterion value(s) : -25.38168
N = 2, M = 5 machine precision = 2.22045e-16
At X0, 0 variables are exactly at the bounds
At iterate 0 f= 25.382 |proj g|= 0.19431
At iterate 1 f = 25.027 |proj g|= 0.13259
At iterate 2 f = 25.014 |proj g|= 1.6725
At iterate 3 f = 25.002 |proj g|= 0.15969
At iterate 4 f = 25.001 |proj g|= 0.17792
At iterate 5 f = 24.999 |proj g|= 0.31318
At iterate 6 f = 24.998 |proj g|= 0.14968
At iterate 7 f = 24.998 |proj g|= 0.03446
At iterate 8 f = 24.998 |proj g|= 0.03458
At iterate 9 f = 24.998 |proj g|= 0.0084816
At iterate 10 f = 24.998 |proj g|= 0.038393
At iterate 11 f = 24.997 |proj g|= 1.3196
At iterate 12 f = 24.997 |proj g|= 1.3327
At iterate 13 f = 24.994 |proj g|= 1.8077
At iterate 14 f = 24.991 |proj g|= 1.8106
At iterate 15 f = 24.975 |proj g|= 1.8136
At iterate 16 f = 24.937 |proj g|= 1.8202
At iterate 17 f = 24.816 |proj g|= 1.8136
At iterate 18 f = 24.652 |proj g|= 0.81261
At iterate 19 f = 24.652 |proj g|= 0.25743
At iterate 20 f = 24.651 |proj g|= 0.0033442
At iterate 21 f = 24.651 |proj g|= 1.4045e-05
iterations 21
function evaluations 30
segments explored during Cauchy searches 22
BFGS updates skipped 0
active bounds at final generalized Cauchy point 1
norm of the final projected gradient 1.40447e-05
final function value 24.6515
F = 24.6515
final value 24.651471
converged
>
> #the points where we want to compute prediction (if a point new.x is added to the doe)
> n.grid <- 20 #you can run it with 100
> x.grid <- y.grid <- seq(0,1,length=n.grid)
> newdata <- expand.grid(x.grid,y.grid)
> precalc.data <- precomputeUpdateData(model=model,integration.points=newdata)
>
> #now we can compute very quickly kriging covariances
> #between these data and any other points
> other.x <- matrix(c(0.6,0.6),ncol=2)
> pred <- predict_nobias_km(object=model,newdata=other.x,type="UK",se.compute=TRUE)
>
> kn <- computeQuickKrigcov(model=model,integration.points=newdata,X.new=other.x,
+ precalc.data=precalc.data,F.newdata=pred$F.newdata,
+ c.newdata=pred$c)
>
> z.grid <- matrix(kn, n.grid, n.grid)
>
> #plots: contour of the criterion, doe points and new point
> image(x=x.grid,y=y.grid,z=z.grid,col=grey.colors(10))
> contour(x=x.grid,y=y.grid,z=z.grid,15,add=TRUE)
> contour(x=x.grid,y=y.grid,z=z.grid,levels=0,add=TRUE,col="blue",lwd=5)
> points(design, col="black", pch=17, lwd=4,cex=2)
> points(other.x, col="red", pch=17, lwd=4,cex=3)
> title("Kriging covariances with the point (0.6,0.6), in red")
>
>
>
>
>
> dev.off()
null device
1
>
|