Supported by Dr. Osamu Ogasawara and  providing providing  . . |
|
Last data update: 2014.03.03 |
Prints a measure of uncertainty for a function of any dimension.DescriptionThis function prints in the whole input domain the value of a given measure of uncertainty.
Possible measures are Usageprint_uncertainty(model, T, type = "pn", ...) Arguments
Valuethe integrated uncertainty Author(s)Clement Chevalier (IMSV, Switzerland, and IRSN, France) ReferencesBect J., Ginsbourger D., Li L., Picheny V., Vazquez E. (2010), Sequential design of computer experiments for the estimation of a probability of failure, Statistics and Computing, pp.1-21, 2011, http://arxiv.org/abs/1009.5177 See Also
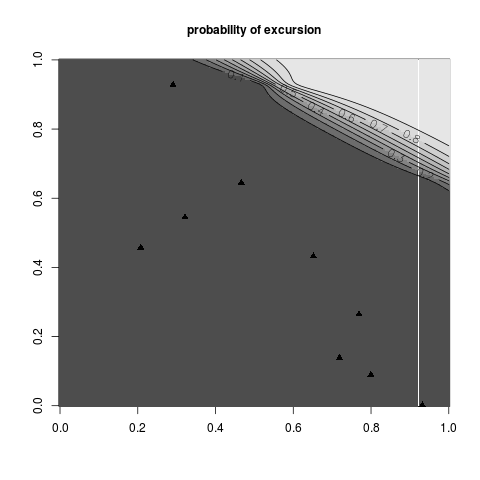
Examples#print_uncertainty set.seed(8) N <- 9 #number of observations T <- 80 #threshold testfun <- branin lower <- c(0,0) upper <- c(1,1) #a 9 points initial design design <- data.frame( matrix(runif(2*N),ncol=2) ) response <- testfun(design) #km object with matern3_2 covariance #params estimated by ML from the observations model <- km(formula=~., design = design, response = response,covtype="matern3_2") #you could do many plots, but only one is run here print_uncertainty(model=model,T=T,main="probability of excursion",type="pn") #print_uncertainty(model=model,T=T,main="Vorob'ev uncertainty",type="vorob") #print_uncertainty(model=model,T=T,main="imse uncertainty",type="imse") #print_uncertainty(model=model,T=T,main="timse uncertainty",type="timse") #print_uncertainty(model=model,T=T,main="sur uncertainty",type="sur") #print_uncertainty(model=model,T=T,main="probability of excursion",type="pn", #vorobmean=TRUE) Results
R version 3.3.1 (2016-06-21) -- "Bug in Your Hair"
Copyright (C) 2016 The R Foundation for Statistical Computing
Platform: x86_64-pc-linux-gnu (64-bit)
R is free software and comes with ABSOLUTELY NO WARRANTY.
You are welcome to redistribute it under certain conditions.
Type 'license()' or 'licence()' for distribution details.
R is a collaborative project with many contributors.
Type 'contributors()' for more information and
'citation()' on how to cite R or R packages in publications.
Type 'demo()' for some demos, 'help()' for on-line help, or
'help.start()' for an HTML browser interface to help.
Type 'q()' to quit R.
> library(KrigInv)
Loading required package: DiceKriging
Loading required package: pbivnorm
Loading required package: rgenoud
## rgenoud (Version 5.7-12.4, Build Date: 2015-07-19)
## See http://sekhon.berkeley.edu/rgenoud for additional documentation.
## Please cite software as:
## Walter Mebane, Jr. and Jasjeet S. Sekhon. 2011.
## ``Genetic Optimization Using Derivatives: The rgenoud package for R.''
## Journal of Statistical Software, 42(11): 1-26.
##
Loading required package: randtoolbox
Loading required package: rngWELL
This is randtoolbox. For overview, type 'help("randtoolbox")'.
> png(filename="/home/ddbj/snapshot/RGM3/R_CC/result/KrigInv/print_uncertainty.Rd_%03d_medium.png", width=480, height=480)
> ### Name: print_uncertainty
> ### Title: Prints a measure of uncertainty for a function of any dimension.
> ### Aliases: print_uncertainty
>
> ### ** Examples
>
> #print_uncertainty
>
> set.seed(8)
> N <- 9 #number of observations
> T <- 80 #threshold
> testfun <- branin
> lower <- c(0,0)
> upper <- c(1,1)
>
> #a 9 points initial design
> design <- data.frame( matrix(runif(2*N),ncol=2) )
> response <- testfun(design)
>
> #km object with matern3_2 covariance
> #params estimated by ML from the observations
> model <- km(formula=~., design = design,
+ response = response,covtype="matern3_2")
optimisation start
------------------
* estimation method : MLE
* optimisation method : BFGS
* analytical gradient : used
* trend model : ~X1 + X2
* covariance model :
- type : matern3_2
- nugget : NO
- parameters lower bounds : 1e-10 1e-10
- parameters upper bounds : 1.448893 1.853021
- best initial criterion value(s) : -25.38168
N = 2, M = 5 machine precision = 2.22045e-16
At X0, 0 variables are exactly at the bounds
At iterate 0 f= 25.382 |proj g|= 0.19431
At iterate 1 f = 25.027 |proj g|= 0.13259
At iterate 2 f = 25.014 |proj g|= 1.6725
At iterate 3 f = 25.002 |proj g|= 0.15969
At iterate 4 f = 25.001 |proj g|= 0.17792
At iterate 5 f = 24.999 |proj g|= 0.31318
At iterate 6 f = 24.998 |proj g|= 0.14968
At iterate 7 f = 24.998 |proj g|= 0.03446
At iterate 8 f = 24.998 |proj g|= 0.03458
At iterate 9 f = 24.998 |proj g|= 0.0084816
At iterate 10 f = 24.998 |proj g|= 0.038393
At iterate 11 f = 24.997 |proj g|= 1.3196
At iterate 12 f = 24.997 |proj g|= 1.3327
At iterate 13 f = 24.994 |proj g|= 1.8077
At iterate 14 f = 24.991 |proj g|= 1.8106
At iterate 15 f = 24.975 |proj g|= 1.8136
At iterate 16 f = 24.937 |proj g|= 1.8202
At iterate 17 f = 24.816 |proj g|= 1.8136
At iterate 18 f = 24.652 |proj g|= 0.81261
At iterate 19 f = 24.652 |proj g|= 0.25743
At iterate 20 f = 24.651 |proj g|= 0.0033442
At iterate 21 f = 24.651 |proj g|= 1.4045e-05
iterations 21
function evaluations 30
segments explored during Cauchy searches 22
BFGS updates skipped 0
active bounds at final generalized Cauchy point 1
norm of the final projected gradient 1.40447e-05
final function value 24.6515
F = 24.6515
final value 24.651471
converged
>
> #you could do many plots, but only one is run here
> print_uncertainty(model=model,T=T,main="probability of excursion",type="pn")
[1] 0.09380482
> #print_uncertainty(model=model,T=T,main="Vorob'ev uncertainty",type="vorob")
> #print_uncertainty(model=model,T=T,main="imse uncertainty",type="imse")
> #print_uncertainty(model=model,T=T,main="timse uncertainty",type="timse")
> #print_uncertainty(model=model,T=T,main="sur uncertainty",type="sur")
> #print_uncertainty(model=model,T=T,main="probability of excursion",type="pn",
> #vorobmean=TRUE)
>
>
>
>
>
> dev.off()
null device
1
>
|