Supported by Dr. Osamu Ogasawara and  providing providing  . . |
|
Last data update: 2014.03.03 |
Logic RegressionDescriptionFit one or a series of Logic Regression models, carry out cross-validation or permutation tests for such models, or fit Monte Carlo Logic Regression models. Logic regression is a (generalized) regression methodology that is
primarily applied when most of the covariates in the data to be
analyzed are binary. The goal of logic regression is to find
predictors that are Boolean (logical) combinations of the original
predictors. Currently the Logic Regression methodology has scoring
functions for linear regression (residual sum of squares), logistic
regression (deviance), classification (misclassification),
proportional hazards models (partial likelihood),
and exponential survival models (log-likelihood). A feature of the
Logic Regression methodology is that it is easily possible to extend
the method to write ones own scoring function if you have a different
scoring function. Usagelogreg(resp, bin, sep, wgt, cens, type, select, ntrees, nleaves,
penalty, seed, kfold, nrep, oldfit, anneal.control, tree.control,
mc.control)
Arguments
DetailsLogic Regression is an adaptive regression methodology that attempts to construct predictors as Boolean combinations of binary covariates. In most regression problems a model is developed that only relates the main effects (the predictors or transformations thereof) to the response. Although interactions between predictors are considered sometimes as well, those interactions are usually kept simple (two- to three-way interactions at most). But often, especially when all predictors are binary, the interaction between many predictors is what causes the differences in response. This issue often arises in the analysis of SNP microarray data or in data mining problems. Given a set of binary predictors X, we try to create new, better predictors for the response by considering combinations of those binary predictors. For example, if the response is binary as well (which is not required in general), we attempt to find decision rules such as “if X1, X2, X3 and X4 are true”, or “X5 or X6 but not X7 are true”, then the response is more likely to be in class 0. In other words, we try to find Boolean statements involving the binary predictors that enhance the prediction for the response. In more specific terms: Let X1,...,Xk be binary predictors, and let Y be a response variable. We try to fit regression models of the form g(E[Y]) = b0 + b1 L1+ ...+ bn Ln, where Lj is a Boolean expression of the predictors X, such as Lj=[(X2 or X4c) and X7]. The above framework includes many forms of regression, such as linear regression (g(E[Y])=E[Y]) and logistic regression (g(E[Y])=log(E[Y]/(1-E[Y]))). For every model type, we define a score function that reflects the “quality” of the model under consideration. For example, for linear regression the score could be the residual sum of squares and for logistic regression the score could be the deviance. We try to find the Boolean expressions in the regression model that minimize the scoring function associated with this model type, estimating the parameters bj simultaneously with the Boolean expressions Lj. In general, any type of model can be considered, as long as a scoring function can be defined. For example, we also implemented the Cox proportional hazards model, using the partial likelihood as the score. Since the number of possible Logic Models we can construct for a given set of predictors is huge, we have to rely on some search algorithms to help us find the best scoring models. We define the move set by a set of standard operations such as splitting and pruning the tree (similar to the terminology introduced by Breiman et al (1984) for CART). We investigated two types of algorithms: a greedy and a simulated annealing algorithm. While the greedy algorithm is very fast, it does not always find a good scoring model. The simulated annealing algorithm usually does, but computationally it is more expensive. Since we have to be certain to find good scoring models, we usually carry out simulated annealing for our case studies. However, as usual, the best scoring model generally over-fits the data, and methods to separate signal and noise are needed. To assess the over-fitting of large models, we offer the option to fit a model of a specific size. For the model selection itself we developed and implemented permutation tests and tests using cross-validation. If sufficient data is available, an analysis using a training and a test set can also be carried out. These tests are rather complicated, so we will not go into detail here and refer you to Ruczinski I, Kooperberg C, LeBlanc ML (2003), cited below. There are two alternatives to the simulated annealing algorithm. One
is a stepwise greedy selection of models. This is when setting
The second alternative is to run a Markov Chain Monte Carlo (MCMC) algorithm.
This is what is done in Monte Carlo Logic Regression. The algorithm used
is a reversible jump MCMC algorithm, due to Green (1995). Other than
the length of the Markov chain, the only parameter that needs to be set
is a parameter for the geometric prior on model size. Other than in many
MCMC problems, the goal in Monte Carlo Logic Regression is not to yield
one single best predicting model, but rather to provide summaries of
all models. These are exactly the elements that are shown above as
the output when MONITORING The help file for Find the best scoring model of any size During the iterations the following information is printed out:
log-temp: This information can be used to judge the convergence of the simulated
annealing algorithm, as described in the help file of
Find the best scoring models for various sizes
During the iterations the same information as for find the best scoring model of any size, for each size model considered. Carry out cross-validation for model selection
Information about the simulated annealing as described above can be printed out. Otherwise, during the cross-validation process information like
The first of the four scores is the training-set score on the current validation sample, the second score is the average of all the training-set scores that have been processed for this model size, the third is the test-set score on the current validation sample, and the fourth score is the average of all the test-set scores that have been processed for this model size. Typically we would prefer the model with the lowest test-set score average over all cross-validation steps. Carry out a permutation test to check for signal in the data
Information about the simulated annealing as described above can be printed out. Otherwise, first the score of the model of size 0 (typically only fitting an intercept) and the score of the best model are printed out. Then during the permutation lines like
are printed. Each score is the result of fitting a logic tree model, on data where the response has been permuted. Typically we would believe that there is signal in the data if most permutations have worse (higher) scores than the best model, but we may believe that there is substantial over-fitting going on if these permutation scores are much better (lower) than the score of the model of size 0. Carry out a permutation test for model selection
To be able to run this option, an object of class
are printed. We can compare these scores to the tree of the same size
and the best tree. If the scores are about the same as the one for
the best tree, we think that the “true” model size may be the one
that is being tested, while if the scores are much worse, the true
model size may be larger. The comparison with the model of the same
size suggests us again how much over-fitting may be going on.
Greedy stepwise selection of Logic Regression models
The scores of the best greedy models of each size are printed. Monte Carlo Logic Regression
A status line is printed every so many iterations. This information is probably not very useful, other than that it helps you figure out how far the code is. PARAMETERS As Logic Regression is written in Fortran 77 some parameters had to be hard coded in. The default values of these parameters are maximum sample size: 20000 If these parameters are not large enough (an error message will let you know this), you need to reset them in slogic.f and recompile. In particular, the statements defining these parameters are
The unfortunate thing is that you will have to change these parameter definitions several times in the code. So search until you have found them all. ValueAn object of the class If an object of class If
If
If
If
If
Author(s)Ingo Ruczinski ingo@jhu.edu and Charles Kooperberg clk@fredhutch.org. ReferencesRuczinski I, Kooperberg C, LeBlanc ML (2003). Logic Regression, Journal of Computational and Graphical Statistics, 12, 475-511. Ruczinski I, Kooperberg C, LeBlanc ML (2002). Logic Regression - methods and software. Proceedings of the MSRI workshop on Nonlinear Estimation and Classification (Eds: D. Denison, M. Hansen, C. Holmes, B. Mallick, B. Yu), Springer: New York, 333-344. Kooperberg C, Ruczinski I, LeBlanc ML, Hsu L (2001). Sequence Analysis using Logic Regression, Genetic Epidemiology, 21, S626-S631. Kooperberg C, Ruczinki I (2005). Identifying interacting SNPs using Monte Carlo Logic Regression, Genetic Epidemiology, 28, 157-170. Selected chapters from the dissertation of Ingo Ruczinski, available from http://kooperberg.fhcrc.org/logic/documents/ingophd-logic.pdf See Also
Examples
data(logreg.savefit1,logreg.savefit2,logreg.savefit3,logreg.savefit4,
logreg.savefit5,logreg.savefit6,logreg.savefit7,logreg.testdat)
myanneal <- logreg.anneal.control(start = -1, end = -4, iter = 500, update = 100)
# in practie we would use 25000 iterations or far more - the use of 500 is only
# to have the examples run fast
## Not run: myanneal <- logreg.anneal.control(start = -1, end = -4, iter = 25000, update = 500)
fit1 <- logreg(resp = logreg.testdat[,1], bin=logreg.testdat[, 2:21], type = 2,
select = 1, ntrees = 2, anneal.control = myanneal)
# the best score should be in the 0.95-1.10 range
plot(fit1)
# you'll probably see X1-X4 as well as a few noise predictors
# use logreg.savefit1 for the results with 25000 iterations
plot(logreg.savefit1)
print(logreg.savefit1)
z <- predict(logreg.savefit1)
plot(z, logreg.testdat[,1]-z, xlab="fitted values", ylab="residuals")
# there are some streaks, thanks to the very discrete predictions
#
# a bit less output
myanneal2 <- logreg.anneal.control(start = -1, end = -4, iter = 500, update = 0)
# in practie we would use 25000 iterations or more - the use of 500 is only
# to have the examples run fast
## Not run: myanneal2 <- logreg.anneal.control(start = -1, end = -4, iter = 25000, update = 0)
#
# fit multiple models
fit2 <- logreg(resp = logreg.testdat[,1], bin=logreg.testdat[, 2:21], type = 2,
select = 2, ntrees = c(1,2), nleaves =c(1,7), anneal.control = myanneal2)
# equivalent
fit2 <- logreg(select = 2, ntrees = c(1,2), nleaves =c(1,7), oldfit = fit1,
anneal.control = myanneal2)
plot(fit2)
# use logreg.savefit2 for the results with 25000 iterations
plot(logreg.savefit2)
print(logreg.savefit2)
# After an initial steep decline, the scores only get slightly better
# for models with more than four leaves and two trees.
#
# cross validation
fit3 <- logreg(resp = logreg.testdat[,1], bin=logreg.testdat[, 2:21], type = 2,
select = 3, ntrees = c(1,2), nleaves=c(1,7), anneal.control = myanneal2)
# equivalent
fit3 <- logreg(select = 3, oldfit = fit2)
plot(fit3)
# use logreg.savefit3 for the results with 25000 iterations
plot(logreg.savefit3)
# 4 leaves, 2 trees should top
# null model test
fit4 <- logreg(resp = logreg.testdat[,1], bin=logreg.testdat[, 2:21], type = 2,
select = 4, ntrees = 2, anneal.control = myanneal2)
# equivalent
fit4 <- logreg(select = 4, anneal.control = myanneal2, oldfit = fit1)
plot(fit4)
# use logreg.savefit4 for the results with 25000 iterations
plot(logreg.savefit4)
# A histogram of the 25 scores obtained from the permutation test. Also shown
# are the scores for the best scoring model with one logic tree, and the null
# model (no tree). Since the permutation scores are not even close to the score
# of the best model with one tree (fit on the original data), there is overwhelming
# evidence against the null hypothesis that there was no signal in the data.
fit5 <- logreg(resp = logreg.testdat[,1], bin=logreg.testdat[, 2:21], type = 2,
select = 5, ntrees = c(1,2), nleaves=c(1,7), anneal.control = myanneal2,
nrep = 10, oldfit = fit2)
# equivalent
fit5 <- logreg(select = 5, nrep = 10, oldfit = fit2)
plot(fit5)
# use logreg.savefit5 for the results with 25000 iterations and 25 permutations
plot(logreg.savefit5)
# The permutation scores improve until we condition on a model with two trees and
# four leaves, and then do not change very much anymore. This indicates that the
# best model has indeed four leaves.
#
# greedy selection
fit6 <- logreg(select = 6, ntrees = 2, nleaves =c(1,12), oldfit = fit1)
plot(fit6)
# use logreg.savefit6 for the results with 25000 iterations
plot(logreg.savefit6)
#
# Monte Carlo Logic Regression
fit7 <- logreg(select = 7, oldfit = fit1, mc.control=
logreg.mc.control(nburn=500, niter=2500, hyperpars=log(2)))
# we need many more iterations for reasonable results
## Not run: logreg.savefit7 <- logreg(select = 7, oldfit = fit1, mc.control=
logreg.mc.control(nburn=1000, niter=100000, hyperpars=log(2)))
## End(Not run)
#
plot(fit7)
# use logreg.savefit7 for the results with 25000 iterations
plot(logreg.savefit7)
Results
R version 3.3.1 (2016-06-21) -- "Bug in Your Hair"
Copyright (C) 2016 The R Foundation for Statistical Computing
Platform: x86_64-pc-linux-gnu (64-bit)
R is free software and comes with ABSOLUTELY NO WARRANTY.
You are welcome to redistribute it under certain conditions.
Type 'license()' or 'licence()' for distribution details.
R is a collaborative project with many contributors.
Type 'contributors()' for more information and
'citation()' on how to cite R or R packages in publications.
Type 'demo()' for some demos, 'help()' for on-line help, or
'help.start()' for an HTML browser interface to help.
Type 'q()' to quit R.
> library(LogicReg)
Loading required package: survival
> png(filename="/home/ddbj/snapshot/RGM3/R_CC/result/LogicReg/logreg.Rd_%03d_medium.png", width=480, height=480)
> ### Name: logreg
> ### Title: Logic Regression
> ### Aliases: logreg
> ### Keywords: logic methods nonparametric tree
>
> ### ** Examples
>
> data(logreg.savefit1,logreg.savefit2,logreg.savefit3,logreg.savefit4,
+ logreg.savefit5,logreg.savefit6,logreg.savefit7,logreg.testdat)
>
> myanneal <- logreg.anneal.control(start = -1, end = -4, iter = 500, update = 100)
> # in practie we would use 25000 iterations or far more - the use of 500 is only
> # to have the examples run fast
> ## Not run: myanneal <- logreg.anneal.control(start = -1, end = -4, iter = 25000, update = 500)
> fit1 <- logreg(resp = logreg.testdat[,1], bin=logreg.testdat[, 2:21], type = 2,
+ select = 1, ntrees = 2, anneal.control = myanneal)
log-temp current score best score acc / rej /sing current parameters
-1.000 1.4941 1.4941 0( 0) 0 0 2.886 0.000 0.000
-1.600 1.2526 1.2261 50( 2) 48 0 2.113 -0.237 1.652
-2.200 1.1005 1.1005 41( 4) 55 0 2.805 1.202 -0.928
-2.800 1.0491 1.0434 22( 4) 74 0 1.868 2.131 -0.063
-3.400 1.0444 1.0434 10( 4) 86 0 1.857 2.120 0.647
-4.000 1.0433 1.0433 2( 4) 94 0 1.852 2.116 0.564
> # the best score should be in the 0.95-1.10 range
> plot(fit1)
> # you'll probably see X1-X4 as well as a few noise predictors
> # use logreg.savefit1 for the results with 25000 iterations
> plot(logreg.savefit1)
> print(logreg.savefit1)
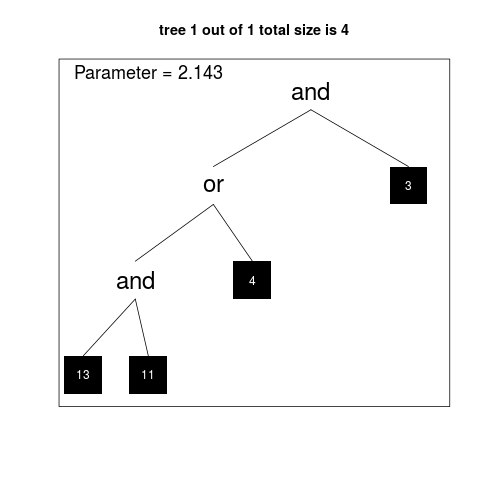
score 0.966
1.98 -1.3 * (((X14 or (not X5)) and ((not X1) and (not X2))) and (((not X3) or X1) or ((not X20) and (not X2)))) +2.15 * (((not X4) or ((not X13) and (not X11))) and (not X3))
> z <- predict(logreg.savefit1)
> plot(z, logreg.testdat[,1]-z, xlab="fitted values", ylab="residuals")
> # there are some streaks, thanks to the very discrete predictions
> #
> # a bit less output
> myanneal2 <- logreg.anneal.control(start = -1, end = -4, iter = 500, update = 0)
> # in practie we would use 25000 iterations or more - the use of 500 is only
> # to have the examples run fast
> ## Not run: myanneal2 <- logreg.anneal.control(start = -1, end = -4, iter = 25000, update = 0)
> #
> # fit multiple models
> fit2 <- logreg(resp = logreg.testdat[,1], bin=logreg.testdat[, 2:21], type = 2,
+ select = 2, ntrees = c(1,2), nleaves =c(1,7), anneal.control = myanneal2)
The number of trees in these models is 1
The model size is 1
The best model with 1 trees of size 1 has a score of 1.1500
The model size is 2
The best model with 1 trees of size 2 has a score of 1.0481
The model size is 3
The best model with 1 trees of size 3 has a score of 1.0481
The model size is 4
The best model with 1 trees of size 4 has a score of 1.0481
The model size is 5
The best model with 1 trees of size 5 has a score of 1.0457
The model size is 6
The best model with 1 trees of size 6 has a score of 1.0481
The model size is 7
The best model with 1 trees of size 7 has a score of 1.0481
The number of trees in these models is 2
The size for this model is smaller than the number of trees you requested.
( 1 versus 2)
To save CPU time, we will skip this run.
On to the next model...
The model size is 2
The best model with 2 trees of size 2 has a score of 1.1168
The model size is 3
The best model with 2 trees of size 3 has a score of 1.0333
The model size is 4
The best model with 2 trees of size 4 has a score of 1.0451
The model size is 5
The best model with 2 trees of size 5 has a score of 1.0309
The model size is 6
The best model with 2 trees of size 6 has a score of 1.0422
The model size is 7
The best model with 2 trees of size 7 has a score of 1.0333
> # equivalent
> fit2 <- logreg(select = 2, ntrees = c(1,2), nleaves =c(1,7), oldfit = fit1,
+ anneal.control = myanneal2)
The number of trees in these models is 1
The model size is 1
The best model with 1 trees of size 1 has a score of 1.1500
The model size is 2
The best model with 1 trees of size 2 has a score of 1.0481
The model size is 3
The best model with 1 trees of size 3 has a score of 1.0481
The model size is 4
The best model with 1 trees of size 4 has a score of 1.0481
The model size is 5
The best model with 1 trees of size 5 has a score of 1.0481
The model size is 6
The best model with 1 trees of size 6 has a score of 1.0437
The model size is 7
The best model with 1 trees of size 7 has a score of 1.1500
The number of trees in these models is 2
The size for this model is smaller than the number of trees you requested.
( 1 versus 2)
To save CPU time, we will skip this run.
On to the next model...
The model size is 2
The best model with 2 trees of size 2 has a score of 1.1168
The model size is 3
The best model with 2 trees of size 3 has a score of 1.0333
The model size is 4
The best model with 2 trees of size 4 has a score of 1.1168
The model size is 5
The best model with 2 trees of size 5 has a score of 1.0454
The model size is 6
The best model with 2 trees of size 6 has a score of 0.9885
The model size is 7
The best model with 2 trees of size 7 has a score of 1.0928
> plot(fit2)
> # use logreg.savefit2 for the results with 25000 iterations
> plot(logreg.savefit2)
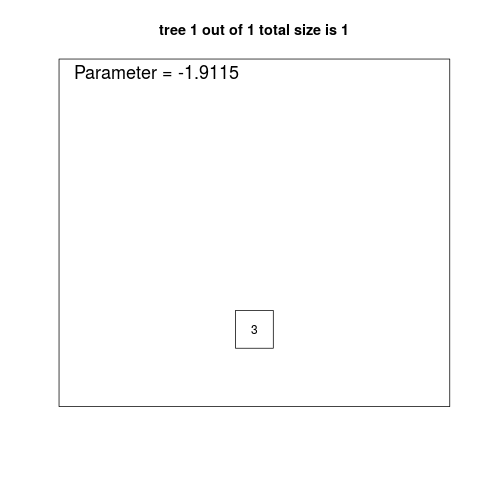
> print(logreg.savefit2)
1 trees with 1 leaves: score is 1.15
3.79 -1.91 * X3
1 trees with 2 leaves: score is 1.048
1.87 +2.13 * ((not X3) and (not X4))
1 trees with 3 leaves: score is 1.046
1.85 +2.14 * ((not X3) and ((not X13) or (not X4)))
1 trees with 4 leaves: score is 1.042
1.86 +2.14 * ((((not X13) and (not X11)) or (not X4)) and (not X3))
1 trees with 5 leaves: score is 1.042
1.86 +2.14 * ((not X3) and ((not X4) or ((not X13) and (not X11))))
1 trees with 6 leaves: score is 1.042
4 -2.14 * (X3 or (X4 and (X13 or (not X19))))
1 trees with 7 leaves: score is 1.04
1.84 +2.15 * ((((not X4) or (not X13)) and (not X3)) or (((not X6) and X14) and ((not X1) and (not X12))))
2 trees with 2 leaves: score is 1.117
1.98 +1.89 * (not X3) -0.904 * X4
2 trees with 3 leaves: score is 1.033
1.58 +0.401 * X1 +2.13 * ((not X3) and (not X4))
2 trees with 4 leaves: score is 0.988
4.12 -1.11 * ((not X1) and (not X2)) -2.12 * (X3 or X4)
2 trees with 5 leaves: score is 0.982
0.77 +1.22 * ((X2 or X1) or X20) +2.12 * ((not X4) and (not X3))
2 trees with 6 leaves: score is 0.979
0.764 +2.13 * ((not X3) and (not X4)) +1.23 * ((X2 or X1) or (X20 and X3))
2 trees with 7 leaves: score is 0.978
1.99 +2.13 * ((not X3) and (not X4)) -1.29 * (((not X7) or ((not X5) or (not X15))) and ((not X1) and (not X2)))
> # After an initial steep decline, the scores only get slightly better
> # for models with more than four leaves and two trees.
> #
> # cross validation
> fit3 <- logreg(resp = logreg.testdat[,1], bin=logreg.testdat[, 2:21], type = 2,
+ select = 3, ntrees = c(1,2), nleaves=c(1,7), anneal.control = myanneal2)
The number of trees in these models is 1
The model size is 1
training-now training-ave test-now test-ave
Step 1 of 10 [ 1 trees; 1 leaves] CV score: 1.1591 1.1591 1.0878 1.0878
Step 2 of 10 [ 1 trees; 1 leaves] CV score: 1.1460 1.1525 1.2104 1.1491
Step 3 of 10 [ 1 trees; 1 leaves] CV score: 1.1181 1.1410 1.4329 1.2437
Step 4 of 10 [ 1 trees; 1 leaves] CV score: 1.1635 1.1467 1.0458 1.1942
Step 5 of 10 [ 1 trees; 1 leaves] CV score: 1.1721 1.1517 0.9495 1.1453
Step 6 of 10 [ 1 trees; 1 leaves] CV score: 1.1344 1.1489 1.3107 1.1729
Step 7 of 10 [ 1 trees; 1 leaves] CV score: 1.1482 1.1488 1.1962 1.1762
Step 8 of 10 [ 1 trees; 1 leaves] CV score: 1.1524 1.1492 1.1532 1.1733
Step 9 of 10 [ 1 trees; 1 leaves] CV score: 1.1648 1.1509 1.0360 1.1581
Step 10 of 10 [ 1 trees; 1 leaves] CV score: 1.1408 1.1499 1.2586 1.1681
The model size is 2
training-now training-ave test-now test-ave
Step 1 of 10 [ 1 trees; 2 leaves] CV score: 1.0468 1.0468 1.0818 1.0818
Step 2 of 10 [ 1 trees; 2 leaves] CV score: 1.0620 1.0544 0.9339 1.0078
Step 3 of 10 [ 1 trees; 2 leaves] CV score: 1.0314 1.0467 1.2123 1.0760
Step 4 of 10 [ 1 trees; 2 leaves] CV score: 1.1635 1.0759 1.0458 1.0684
Step 5 of 10 [ 1 trees; 2 leaves] CV score: 1.0621 1.0732 0.9345 1.0416
Step 6 of 10 [ 1 trees; 2 leaves] CV score: 1.1344 1.0834 1.3107 1.0865
Step 7 of 10 [ 1 trees; 2 leaves] CV score: 1.0530 1.0790 1.0246 1.0776
Step 8 of 10 [ 1 trees; 2 leaves] CV score: 1.0464 1.0750 1.0853 1.0786
Step 9 of 10 [ 1 trees; 2 leaves] CV score: 1.1648 1.0849 1.0360 1.0739
Step 10 of 10 [ 1 trees; 2 leaves] CV score: 1.1408 1.0905 1.2586 1.0923
The model size is 3
training-now training-ave test-now test-ave
Step 1 of 10 [ 1 trees; 3 leaves] CV score: 1.0468 1.0468 1.0818 1.0818
Step 2 of 10 [ 1 trees; 3 leaves] CV score: 1.0585 1.0527 1.0480 1.0649
Step 3 of 10 [ 1 trees; 3 leaves] CV score: 1.0314 1.0456 1.2123 1.1140
Step 4 of 10 [ 1 trees; 3 leaves] CV score: 1.0502 1.0467 1.0511 1.0983
Step 5 of 10 [ 1 trees; 3 leaves] CV score: 1.1721 1.0718 0.9495 1.0685
Step 6 of 10 [ 1 trees; 3 leaves] CV score: 1.1344 1.0822 1.3107 1.1089
Step 7 of 10 [ 1 trees; 3 leaves] CV score: 1.0506 1.0777 1.0224 1.0966
Step 8 of 10 [ 1 trees; 3 leaves] CV score: 1.0464 1.0738 1.0853 1.0951
Step 9 of 10 [ 1 trees; 3 leaves] CV score: 1.0557 1.0718 1.0000 1.0846
Step 10 of 10 [ 1 trees; 3 leaves] CV score: 1.0330 1.0679 1.1805 1.0942
The model size is 4
training-now training-ave test-now test-ave
Step 1 of 10 [ 1 trees; 4 leaves] CV score: 1.1591 1.1591 1.0878 1.0878
Step 2 of 10 [ 1 trees; 4 leaves] CV score: 1.0620 1.1105 0.9339 1.0108
Step 3 of 10 [ 1 trees; 4 leaves] CV score: 1.0314 1.0842 1.2123 1.0780
Step 4 of 10 [ 1 trees; 4 leaves] CV score: 1.0502 1.0757 1.0511 1.0712
Step 5 of 10 [ 1 trees; 4 leaves] CV score: 1.1721 1.0950 0.9495 1.0469
Step 6 of 10 [ 1 trees; 4 leaves] CV score: 1.1344 1.1015 1.3107 1.0909
Step 7 of 10 [ 1 trees; 4 leaves] CV score: 1.1482 1.1082 1.1962 1.1059
Step 8 of 10 [ 1 trees; 4 leaves] CV score: 1.0412 1.0998 1.1072 1.1061
Step 9 of 10 [ 1 trees; 4 leaves] CV score: 1.0557 1.0949 1.0000 1.0943
Step 10 of 10 [ 1 trees; 4 leaves] CV score: 1.1408 1.0995 1.2586 1.1107
The model size is 5
training-now training-ave test-now test-ave
Step 1 of 10 [ 1 trees; 5 leaves] CV score: 1.0468 1.0468 1.0818 1.0818
Step 2 of 10 [ 1 trees; 5 leaves] CV score: 1.0620 1.0544 0.9339 1.0078
Step 3 of 10 [ 1 trees; 5 leaves] CV score: 1.0264 1.0451 1.2116 1.0758
Step 4 of 10 [ 1 trees; 5 leaves] CV score: 1.0502 1.0463 1.0511 1.0696
Step 5 of 10 [ 1 trees; 5 leaves] CV score: 1.1721 1.0715 0.9495 1.0456
Step 6 of 10 [ 1 trees; 5 leaves] CV score: 1.0370 1.0657 1.1682 1.0660
Step 7 of 10 [ 1 trees; 5 leaves] CV score: 1.0530 1.0639 1.0246 1.0601
Step 8 of 10 [ 1 trees; 5 leaves] CV score: 1.0464 1.0617 1.0853 1.0632
Step 9 of 10 [ 1 trees; 5 leaves] CV score: 1.1648 1.0732 1.0360 1.0602
Step 10 of 10 [ 1 trees; 5 leaves] CV score: 1.0330 1.0692 1.1805 1.0722
The model size is 6
training-now training-ave test-now test-ave
Step 1 of 10 [ 1 trees; 6 leaves] CV score: 1.1591 1.1591 1.0878 1.0878
Step 2 of 10 [ 1 trees; 6 leaves] CV score: 1.1429 1.1510 1.2106 1.1492
Step 3 of 10 [ 1 trees; 6 leaves] CV score: 1.1181 1.1400 1.4329 1.2438
Step 4 of 10 [ 1 trees; 6 leaves] CV score: 1.0502 1.1176 1.0511 1.1956
Step 5 of 10 [ 1 trees; 6 leaves] CV score: 1.1694 1.1280 1.0139 1.1592
Step 6 of 10 [ 1 trees; 6 leaves] CV score: 1.0315 1.1119 1.1693 1.1609
Step 7 of 10 [ 1 trees; 6 leaves] CV score: 1.0506 1.1031 1.0224 1.1411
Step 8 of 10 [ 1 trees; 6 leaves] CV score: 1.0412 1.0954 1.1072 1.1369
Step 9 of 10 [ 1 trees; 6 leaves] CV score: 1.0557 1.0910 1.0000 1.1217
Step 10 of 10 [ 1 trees; 6 leaves] CV score: 1.0306 1.0849 1.1804 1.1276
The model size is 7
training-now training-ave test-now test-ave
Step 1 of 10 [ 1 trees; 7 leaves] CV score: 1.0426 1.0426 1.0960 1.0960
Step 2 of 10 [ 1 trees; 7 leaves] CV score: 1.0620 1.0523 1.1272 1.1116
Step 3 of 10 [ 1 trees; 7 leaves] CV score: 1.0287 1.0444 1.2120 1.1451
Step 4 of 10 [ 1 trees; 7 leaves] CV score: 1.1332 1.0666 1.0321 1.1168
Step 5 of 10 [ 1 trees; 7 leaves] CV score: 1.1721 1.0877 0.9495 1.0834
Step 6 of 10 [ 1 trees; 7 leaves] CV score: 1.0370 1.0793 1.1682 1.0975
Step 7 of 10 [ 1 trees; 7 leaves] CV score: 1.1482 1.0891 1.1962 1.1116
Step 8 of 10 [ 1 trees; 7 leaves] CV score: 1.0464 1.0838 1.0853 1.1083
Step 9 of 10 [ 1 trees; 7 leaves] CV score: 1.0532 1.0804 0.9984 1.0961
Step 10 of 10 [ 1 trees; 7 leaves] CV score: 1.0360 1.0759 1.1780 1.1043
The number of trees in these models is 2
The size for this model is smaller than the number of trees you requested.
( 1 versus 2)
To save CPU time, we will skip this run.
On to the next model...
The model size is 2
training-now training-ave test-now test-ave
Step 1 of 10 [ 2 trees; 2 leaves] CV score: 1.1217 1.1217 1.1067 1.1067
Step 2 of 10 [ 2 trees; 2 leaves] CV score: 1.1238 1.1228 1.0981 1.1024
Step 3 of 10 [ 2 trees; 2 leaves] CV score: 1.0895 1.1117 1.3799 1.1949
Step 4 of 10 [ 2 trees; 2 leaves] CV score: 1.1240 1.1148 1.0906 1.1688
Step 5 of 10 [ 2 trees; 2 leaves] CV score: 1.1356 1.1189 0.9641 1.1279
Step 6 of 10 [ 2 trees; 2 leaves] CV score: 1.1073 1.1170 1.2433 1.1471
Step 7 of 10 [ 2 trees; 2 leaves] CV score: 1.1172 1.1170 1.1520 1.1478
Step 8 of 10 [ 2 trees; 2 leaves] CV score: 1.1176 1.1171 1.1472 1.1477
Step 9 of 10 [ 2 trees; 2 leaves] CV score: 1.1244 1.1179 1.0876 1.1411
Step 10 of 10 [ 2 trees; 2 leaves] CV score: 1.1060 1.1167 1.2517 1.1521
The model size is 3
training-now training-ave test-now test-ave
Step 1 of 10 [ 2 trees; 3 leaves] CV score: 1.0302 1.0302 1.0951 1.0951
Step 2 of 10 [ 2 trees; 3 leaves] CV score: 1.0455 1.0379 0.9489 1.0220
Step 3 of 10 [ 2 trees; 3 leaves] CV score: 1.0759 1.0505 1.4647 1.1696
Step 4 of 10 [ 2 trees; 3 leaves] CV score: 1.0382 1.0475 1.0217 1.1326
Step 5 of 10 [ 2 trees; 3 leaves] CV score: 1.0481 1.0476 0.9232 1.0907
Step 6 of 10 [ 2 trees; 3 leaves] CV score: 1.0209 1.0431 1.1765 1.1050
Step 7 of 10 [ 2 trees; 3 leaves] CV score: 1.0398 1.0427 1.0057 1.0908
Step 8 of 10 [ 2 trees; 3 leaves] CV score: 1.1176 1.0520 1.1472 1.0979
Step 9 of 10 [ 2 trees; 3 leaves] CV score: 1.0566 1.0525 1.0070 1.0878
Step 10 of 10 [ 2 trees; 3 leaves] CV score: 1.0371 1.0510 1.1894 1.0979
The model size is 4
training-now training-ave test-now test-ave
Step 1 of 10 [ 2 trees; 4 leaves] CV score: 1.0302 1.0302 1.0951 1.0951
Step 2 of 10 [ 2 trees; 4 leaves] CV score: 0.9992 1.0147 0.9196 1.0074
Step 3 of 10 [ 2 trees; 4 leaves] CV score: 1.0719 1.0338 1.3864 1.1337
Step 4 of 10 [ 2 trees; 4 leaves] CV score: 1.0427 1.0360 1.0611 1.1156
Step 5 of 10 [ 2 trees; 4 leaves] CV score: 1.0554 1.0399 0.9353 1.0795
Step 6 of 10 [ 2 trees; 4 leaves] CV score: 1.1047 1.0507 1.2982 1.1160
Step 7 of 10 [ 2 trees; 4 leaves] CV score: 0.9986 1.0433 0.9227 1.0884
Step 8 of 10 [ 2 trees; 4 leaves] CV score: 1.0242 1.0409 1.0865 1.0881
Step 9 of 10 [ 2 trees; 4 leaves] CV score: 1.0420 1.0410 0.9759 1.0757
Step 10 of 10 [ 2 trees; 4 leaves] CV score: 1.0242 1.0393 1.1519 1.0833
The model size is 5
training-now training-ave test-now test-ave
Step 1 of 10 [ 2 trees; 5 leaves] CV score: 0.9809 0.9809 1.0885 1.0885
Step 2 of 10 [ 2 trees; 5 leaves] CV score: 1.1238 1.0524 1.0981 1.0933
Step 3 of 10 [ 2 trees; 5 leaves] CV score: 1.0109 1.0386 1.2590 1.1485
Step 4 of 10 [ 2 trees; 5 leaves] CV score: 1.1240 1.0599 1.0906 1.1341
Step 5 of 10 [ 2 trees; 5 leaves] CV score: 1.0566 1.0593 0.9524 1.0977
Step 6 of 10 [ 2 trees; 5 leaves] CV score: 0.9749 1.0452 1.1388 1.1046
Step 7 of 10 [ 2 trees; 5 leaves] CV score: 1.1172 1.0555 1.1520 1.1113
Step 8 of 10 [ 2 trees; 5 leaves] CV score: 1.0316 1.0525 1.0989 1.1098
Step 9 of 10 [ 2 trees; 5 leaves] CV score: 0.9919 1.0458 0.9901 1.0965
Step 10 of 10 [ 2 trees; 5 leaves] CV score: 1.1060 1.0518 1.2517 1.1120
The model size is 6
training-now training-ave test-now test-ave
Step 1 of 10 [ 2 trees; 6 leaves] CV score: 1.0192 1.0192 1.1316 1.1316
Step 2 of 10 [ 2 trees; 6 leaves] CV score: 1.0468 1.0330 1.0736 1.1026
Step 3 of 10 [ 2 trees; 6 leaves] CV score: 1.0033 1.0231 1.2631 1.1561
Step 4 of 10 [ 2 trees; 6 leaves] CV score: 0.9947 1.0160 0.9627 1.1078
Step 5 of 10 [ 2 trees; 6 leaves] CV score: 1.0026 1.0133 0.8833 1.0629
Step 6 of 10 [ 2 trees; 6 leaves] CV score: 1.0096 1.0127 1.1927 1.0845
Step 7 of 10 [ 2 trees; 6 leaves] CV score: 1.0195 1.0137 0.9913 1.0712
Step 8 of 10 [ 2 trees; 6 leaves] CV score: 1.0861 1.0227 1.0811 1.0724
Step 9 of 10 [ 2 trees; 6 leaves] CV score: 1.0566 1.0265 1.0070 1.0651
Step 10 of 10 [ 2 trees; 6 leaves] CV score: 1.0198 1.0258 1.1583 1.0745
The model size is 7
training-now training-ave test-now test-ave
Step 1 of 10 [ 2 trees; 7 leaves] CV score: 0.9734 0.9734 1.0944 1.0944
Step 2 of 10 [ 2 trees; 7 leaves] CV score: 1.0388 1.0061 1.1215 1.1079
Step 3 of 10 [ 2 trees; 7 leaves] CV score: 1.0021 1.0048 1.2663 1.1607
Step 4 of 10 [ 2 trees; 7 leaves] CV score: 1.0500 1.0161 1.0774 1.1399
Step 5 of 10 [ 2 trees; 7 leaves] CV score: 1.0629 1.0255 0.9407 1.1000
Step 6 of 10 [ 2 trees; 7 leaves] CV score: 1.0209 1.0247 1.1765 1.1128
Step 7 of 10 [ 2 trees; 7 leaves] CV score: 1.0541 1.0289 1.0316 1.1012
Step 8 of 10 [ 2 trees; 7 leaves] CV score: 1.0994 1.0377 1.1609 1.1087
Step 9 of 10 [ 2 trees; 7 leaves] CV score: 1.1353 1.0486 1.0622 1.1035
Step 10 of 10 [ 2 trees; 7 leaves] CV score: 1.0973 1.0534 1.1866 1.1118
> # equivalent
> fit3 <- logreg(select = 3, oldfit = fit2)
The number of trees in these models is 1
The model size is 1
training-now training-ave test-now test-ave
Step 1 of 10 [ 1 trees; 1 leaves] CV score: 1.0943 1.0943 1.5956 1.5956
Step 2 of 10 [ 1 trees; 1 leaves] CV score: 1.1643 1.1293 1.0350 1.3153
Step 3 of 10 [ 1 trees; 1 leaves] CV score: 1.1710 1.1432 0.9617 1.1975
Step 4 of 10 [ 1 trees; 1 leaves] CV score: 1.1565 1.1465 1.1128 1.1763
Step 5 of 10 [ 1 trees; 1 leaves] CV score: 1.1367 1.1445 1.2927 1.1996
Step 6 of 10 [ 1 trees; 1 leaves] CV score: 1.1724 1.1492 0.9507 1.1581
Step 7 of 10 [ 1 trees; 1 leaves] CV score: 1.1648 1.1514 1.0300 1.1398
Step 8 of 10 [ 1 trees; 1 leaves] CV score: 1.1533 1.1517 1.1463 1.1406
Step 9 of 10 [ 1 trees; 1 leaves] CV score: 1.1492 1.1514 1.1845 1.1455
Step 10 of 10 [ 1 trees; 1 leaves] CV score: 1.1366 1.1499 1.2907 1.1600
The model size is 2
training-now training-ave test-now test-ave
Step 1 of 10 [ 1 trees; 2 leaves] CV score: 1.0053 1.0053 1.4018 1.4018
Step 2 of 10 [ 1 trees; 2 leaves] CV score: 1.1643 1.0848 1.0350 1.2184
Step 3 of 10 [ 1 trees; 2 leaves] CV score: 1.1710 1.1135 0.9617 1.1329
Step 4 of 10 [ 1 trees; 2 leaves] CV score: 1.1565 1.1243 1.1128 1.1278
Step 5 of 10 [ 1 trees; 2 leaves] CV score: 1.1367 1.1267 1.2927 1.1608
Step 6 of 10 [ 1 trees; 2 leaves] CV score: 1.0657 1.1166 0.8936 1.1163
Step 7 of 10 [ 1 trees; 2 leaves] CV score: 1.0664 1.1094 0.8861 1.0834
Step 8 of 10 [ 1 trees; 2 leaves] CV score: 1.1533 1.1149 1.1463 1.0913
Step 9 of 10 [ 1 trees; 2 leaves] CV score: 1.0423 1.1068 1.1262 1.0952
Step 10 of 10 [ 1 trees; 2 leaves] CV score: 1.0166 1.0978 1.3244 1.1181
The model size is 3
training-now training-ave test-now test-ave
Step 1 of 10 [ 1 trees; 3 leaves] CV score: 1.0025 1.0025 1.4023 1.4023
Step 2 of 10 [ 1 trees; 3 leaves] CV score: 1.0499 1.0262 1.0554 1.2288
Step 3 of 10 [ 1 trees; 3 leaves] CV score: 1.1710 1.0744 0.9617 1.1398
Step 4 of 10 [ 1 trees; 3 leaves] CV score: 1.0736 1.0742 0.8031 1.0556
Step 5 of 10 [ 1 trees; 3 leaves] CV score: 1.0466 1.0687 1.0860 1.0617
Step 6 of 10 [ 1 trees; 3 leaves] CV score: 1.0614 1.0675 0.9110 1.0366
Step 7 of 10 [ 1 trees; 3 leaves] CV score: 1.0664 1.0673 0.8861 1.0151
Step 8 of 10 [ 1 trees; 3 leaves] CV score: 1.1533 1.0781 1.1463 1.0315
Step 9 of 10 [ 1 trees; 3 leaves] CV score: 1.0423 1.0741 1.1262 1.0420
Step 10 of 10 [ 1 trees; 3 leaves] CV score: 1.0166 1.0684 1.3244 1.0703
The model size is 4
training-now training-ave test-now test-ave
Step 1 of 10 [ 1 trees; 4 leaves] CV score: 1.0053 1.0053 1.4018 1.4018
Step 2 of 10 [ 1 trees; 4 leaves] CV score: 1.1643 1.0848 1.0350 1.2184
Step 3 of 10 [ 1 trees; 4 leaves] CV score: 1.0665 1.0787 0.8542 1.0970
Step 4 of 10 [ 1 trees; 4 leaves] CV score: 1.1565 1.0981 1.1128 1.1009
Step 5 of 10 [ 1 trees; 4 leaves] CV score: 1.0466 1.0878 1.0860 1.0980
Step 6 of 10 [ 1 trees; 4 leaves] CV score: 1.1724 1.1019 0.9507 1.0734
Step 7 of 10 [ 1 trees; 4 leaves] CV score: 1.0664 1.0969 0.8861 1.0467
Step 8 of 10 [ 1 trees; 4 leaves] CV score: 1.0431 1.0901 1.1162 1.0554
Step 9 of 10 [ 1 trees; 4 leaves] CV score: 1.0392 1.0845 1.1308 1.0637
Step 10 of 10 [ 1 trees; 4 leaves] CV score: 1.0166 1.0777 1.3244 1.0898
The model size is 5
training-now training-ave test-now test-ave
Step 1 of 10 [ 1 trees; 5 leaves] CV score: 1.0053 1.0053 1.4018 1.4018
Step 2 of 10 [ 1 trees; 5 leaves] CV score: 1.0499 1.0276 1.0554 1.2286
Step 3 of 10 [ 1 trees; 5 leaves] CV score: 1.1710 1.0754 0.9617 1.1396
Step 4 of 10 [ 1 trees; 5 leaves] CV score: 1.1565 1.0957 1.1128 1.1329
Step 5 of 10 [ 1 trees; 5 leaves] CV score: 1.1325 1.1030 1.2909 1.1645
Step 6 of 10 [ 1 trees; 5 leaves] CV score: 1.0657 1.0968 0.8936 1.1194
Step 7 of 10 [ 1 trees; 5 leaves] CV score: 1.0664 1.0925 0.8861 1.0861
Step 8 of 10 [ 1 trees; 5 leaves] CV score: 1.0431 1.0863 1.1162 1.0898
Step 9 of 10 [ 1 trees; 5 leaves] CV score: 1.0423 1.0814 1.1262 1.0939
Step 10 of 10 [ 1 trees; 5 leaves] CV score: 1.0166 1.0749 1.3244 1.1169
The model size is 6
training-now training-ave test-now test-ave
Step 1 of 10 [ 1 trees; 6 leaves] CV score: 1.0943 1.0943 1.5956 1.5956
Step 2 of 10 [ 1 trees; 6 leaves] CV score: 1.0499 1.0721 1.0554 1.3255
Step 3 of 10 [ 1 trees; 6 leaves] CV score: 1.0690 1.0711 0.8560 1.1690
Step 4 of 10 [ 1 trees; 6 leaves] CV score: 1.0710 1.0710 0.8021 1.0773
Step 5 of 10 [ 1 trees; 6 leaves] CV score: 1.0443 1.0657 1.0828 1.0784
Step 6 of 10 [ 1 trees; 6 leaves] CV score: 1.0657 1.0657 0.8936 1.0476
Step 7 of 10 [ 1 trees; 6 leaves] CV score: 1.0664 1.0658 0.8861 1.0245
Step 8 of 10 [ 1 trees; 6 leaves] CV score: 1.0431
|