|
Last data update: 2014.03.03
|
R: Microarray probes filtering
Microarray probes filtering
Description
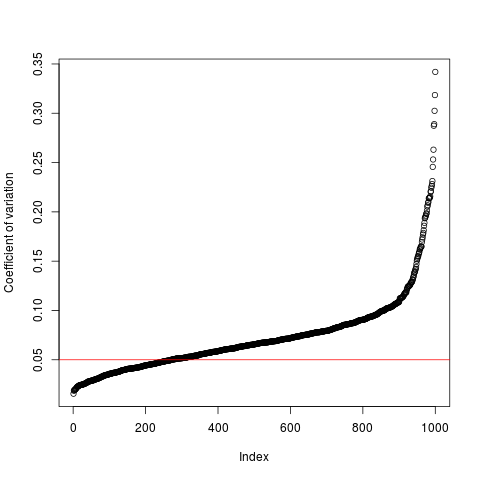
Function to filter microarray probes according to coefficient of variation
Usage
cv.filter(data, cutoff = 0.05)
Arguments
data |
expression matrix with probes in rows and samples in columns
|
cutoff |
cutoff value for filtering
|
Value
Expression matrix, probes with CV below cutoff are filtered out.
Author(s)
Ivana Ihnatova
Examples
data(Singhdata)
data<-Singhdata$esets[[1]][1:1000,]
data.filtered<-cv.filter(data)
head(data.filtered)
Results
R version 3.3.1 (2016-06-21) -- "Bug in Your Hair"
Copyright (C) 2016 The R Foundation for Statistical Computing
Platform: x86_64-pc-linux-gnu (64-bit)
R is free software and comes with ABSOLUTELY NO WARRANTY.
You are welcome to redistribute it under certain conditions.
Type 'license()' or 'licence()' for distribution details.
R is a collaborative project with many contributors.
Type 'contributors()' for more information and
'citation()' on how to cite R or R packages in publications.
Type 'demo()' for some demos, 'help()' for on-line help, or
'help.start()' for an HTML browser interface to help.
Type 'q()' to quit R.
> library(MAMA)
Loading required package: genefilter
Loading required package: metaMA
Attaching package: 'metaMA'
The following object is masked from 'package:genefilter':
rowVars
Loading required package: xtable
Loading required package: multtest
Loading required package: BiocGenerics
Loading required package: parallel
Attaching package: 'BiocGenerics'
The following objects are masked from 'package:parallel':
clusterApply, clusterApplyLB, clusterCall, clusterEvalQ,
clusterExport, clusterMap, parApply, parCapply, parLapply,
parLapplyLB, parRapply, parSapply, parSapplyLB
The following objects are masked from 'package:stats':
IQR, mad, xtabs
The following objects are masked from 'package:base':
Filter, Find, Map, Position, Reduce, anyDuplicated, append,
as.data.frame, cbind, colnames, do.call, duplicated, eval, evalq,
get, grep, grepl, intersect, is.unsorted, lapply, lengths, mapply,
match, mget, order, paste, pmax, pmax.int, pmin, pmin.int, rank,
rbind, rownames, sapply, setdiff, sort, table, tapply, union,
unique, unsplit
Loading required package: Biobase
Welcome to Bioconductor
Vignettes contain introductory material; view with
'browseVignettes()'. To cite Bioconductor, see
'citation("Biobase")', and for packages 'citation("pkgname")'.
Loading required package: gtools
Loading required package: grid
Loading required package: GeneMeta
Attaching package: 'MAMA'
The following objects are masked from 'package:GeneMeta':
multExpFDR, zScoreFDR, zScorePermuted, zScores
> png(filename="/home/ddbj/snapshot/RGM3/R_CC/result/MAMA/cv.filter.Rd_%03d_medium.png", width=480, height=480)
> ### Name: cv.filter
> ### Title: Microarray probes filtering
> ### Aliases: cv.filter
> ### Keywords: manip
>
> ### ** Examples
>
> data(Singhdata)
> data<-Singhdata$esets[[1]][1:1000,]
> data.filtered<-cv.filter(data)
> head(data.filtered)
Normal Normal.1 Normal.2 Normal.3 Normal.4 Normal.5 Normal.6
3 3.783717 4.118151 4.216871 4.223820 4.364816 4.092585 3.999631
4 3.152045 3.633564 3.767186 3.901121 4.001623 3.851386 3.703695
5 5.452305 6.716646 6.431922 6.612021 6.348126 6.762278 6.582096
7 7.207633 5.287455 5.419958 5.413211 5.378318 5.330816 5.280372
8 3.077360 3.383413 3.646509 3.529396 3.436742 3.634367 3.428753
10 6.833431 11.661911 11.549448 11.347327 11.535383 12.120565 11.493540
Normal.7 Normal.8 Normal.9 Tumour Tumour.1 Tumour.2 Tumour.3
3 4.294531 4.254407 4.010793 3.693758 3.837384 3.939096 3.681716
4 3.771112 3.723193 3.507246 3.335860 3.448102 3.423283 3.193589
5 6.648011 6.512724 6.342057 5.623833 6.284547 6.348526 5.872347
7 5.449844 5.343616 5.037288 8.411844 5.092543 5.420076 5.999140
8 3.803338 3.486255 3.453213 3.349204 3.461864 3.514328 3.005631
10 11.950284 11.611832 11.228292 9.235940 11.559250 11.574905 10.786439
Tumour.4 Tumour.5 Tumour.6 Tumour.7 Tumour.8 Tumour.9
3 3.947196 3.861054 3.635040 3.898488 4.311007 3.786629
4 3.536552 3.501316 3.238554 3.311750 3.591361 3.358651
5 6.204390 6.323174 5.756556 6.148963 6.439817 5.940263
7 5.197023 5.916329 8.696407 5.060513 5.389452 6.057525
8 3.287253 3.383124 3.125745 3.429256 3.685988 3.515686
10 11.249121 11.332153 8.763352 11.206492 11.524489 10.477612
>
>
>
>
>
> dev.off()
null device
1
>
|
|
 providing
providing  .
.