Supported by Dr. Osamu Ogasawara and  providing providing  . . |
|
Last data update: 2014.03.03 |
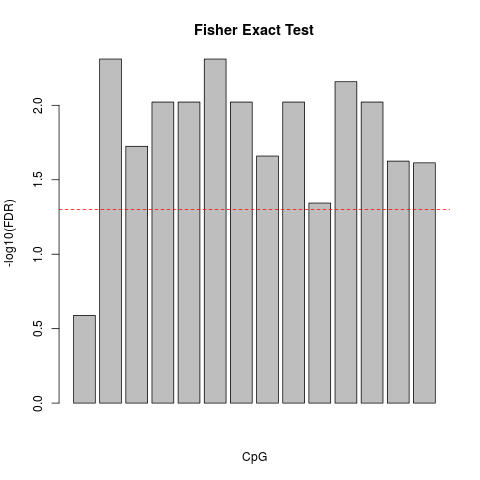
Perform Fisher Exact Test on Methylation Data.DescriptionFisher exact test is a test to calculate the statistical significance using contingency table. It was used to find the statistically significant differences in the methylation status of one particular CpG site between healthy and tumor sample. Contingency matrix was created for each CpG site. P-value was corrected for multiple testing using Benjamini-Hochberg method to calculate False Discovery Rate (FDR) Usagefishertest_cpg(healthy, tumor, plot = TRUE, main = "Fisher Exact Test") Arguments
ValueVector containing p-values. Author(s)Muhammad Ahmer Jamil, Prof. Holger Frohlich, Priv.-Doz. Dr. Osman El-Maarri Maintainer: Muhammad Ahmer Jamil engr.ahmerjamil@gmail.com See Also
Examples
healthy = system.file("extdata", "Healthy.fasta", package = "MethTargetedNGS")
tumor = system.file("extdata", "Tumor.fasta", package = "MethTargetedNGS")
reference = system.file("extdata", "Reference.fasta", package = "MethTargetedNGS")
healthy = methAlign(healthy,reference)
tumor = methAlign(tumor,reference)
fisherexacttest <- fishertest_cpg(healthy,tumor)
Results
R version 3.3.1 (2016-06-21) -- "Bug in Your Hair"
Copyright (C) 2016 The R Foundation for Statistical Computing
Platform: x86_64-pc-linux-gnu (64-bit)
R is free software and comes with ABSOLUTELY NO WARRANTY.
You are welcome to redistribute it under certain conditions.
Type 'license()' or 'licence()' for distribution details.
R is a collaborative project with many contributors.
Type 'contributors()' for more information and
'citation()' on how to cite R or R packages in publications.
Type 'demo()' for some demos, 'help()' for on-line help, or
'help.start()' for an HTML browser interface to help.
Type 'q()' to quit R.
> library(MethTargetedNGS)
Loading required package: stringr
Loading required package: seqinr
Loading required package: gplots
Attaching package: 'gplots'
The following object is masked from 'package:stats':
lowess
Loading required package: Biostrings
Loading required package: BiocGenerics
Loading required package: parallel
Attaching package: 'BiocGenerics'
The following objects are masked from 'package:parallel':
clusterApply, clusterApplyLB, clusterCall, clusterEvalQ,
clusterExport, clusterMap, parApply, parCapply, parLapply,
parLapplyLB, parRapply, parSapply, parSapplyLB
The following objects are masked from 'package:stats':
IQR, mad, xtabs
The following objects are masked from 'package:base':
Filter, Find, Map, Position, Reduce, anyDuplicated, append,
as.data.frame, cbind, colnames, do.call, duplicated, eval, evalq,
get, grep, grepl, intersect, is.unsorted, lapply, lengths, mapply,
match, mget, order, paste, pmax, pmax.int, pmin, pmin.int, rank,
rbind, rownames, sapply, setdiff, sort, table, tapply, union,
unique, unsplit
Loading required package: S4Vectors
Loading required package: stats4
Attaching package: 'S4Vectors'
The following object is masked from 'package:gplots':
space
The following objects are masked from 'package:base':
colMeans, colSums, expand.grid, rowMeans, rowSums
Loading required package: IRanges
Loading required package: XVector
Attaching package: 'Biostrings'
The following object is masked from 'package:seqinr':
translate
> png(filename="/home/ddbj/snapshot/RGM3/R_BC/result/MethTargetedNGS/fishertest_cpg.Rd_%03d_medium.png", width=480, height=480)
> ### Name: fishertest_cpg
> ### Title: Perform Fisher Exact Test on Methylation Data.
> ### Aliases: fishertest_cpg
> ### Keywords: Significant CpG Methylation Fisher Test
>
> ### ** Examples
>
> healthy = system.file("extdata", "Healthy.fasta", package = "MethTargetedNGS")
> tumor = system.file("extdata", "Tumor.fasta", package = "MethTargetedNGS")
> reference = system.file("extdata", "Reference.fasta", package = "MethTargetedNGS")
>
> healthy = methAlign(healthy,reference)
Time difference of 2.15 secs
> tumor = methAlign(tumor,reference)
Time difference of 1.81 secs
> fisherexacttest <- fishertest_cpg(healthy,tumor)
>
>
>
>
>
> dev.off()
null device
1
>
|