|
Last data update: 2014.03.03
|
R: Plot univariate local false discovery output
| plot.fdr1d.result | R Documentation |
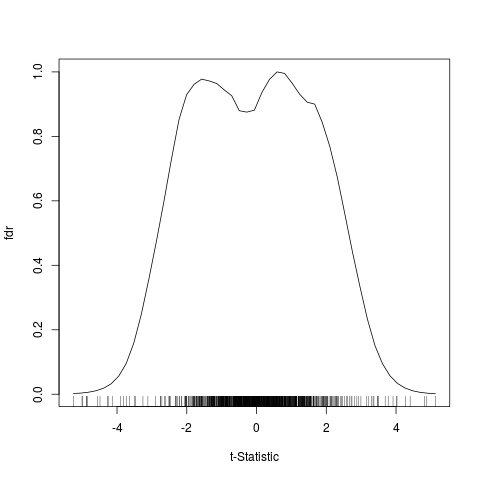
Plot univariate local false discovery output
Description
A plotting function for fdr1d.
Usage
plot.fdr1d.result(x, add = FALSE, grid = FALSE, rug = TRUE,
xlab = "t-Statistic", ylab = "fdr", lcol = "black", ...)
Arguments
x |
output from fdr1d
|
add |
logical value indicating whether to create a new plot or add to an existing one
|
grid |
logical value indicating whether to show the intervals used for calculating the fdr.
|
rug |
logical value indicating whether to add a 1D scatterplot showing the observed test statistics
|
xlab, ylab |
the usual axis labels
|
lcol |
the color of the lines
|
... |
extra options passed to plot.default.
|
Author(s)
A Ploner
See Also
fdr1d
Examples
example(fdr1d)
plot(res1d, grid=TRUE, rug=FALSE)
Results
R version 3.3.1 (2016-06-21) -- "Bug in Your Hair"
Copyright (C) 2016 The R Foundation for Statistical Computing
Platform: x86_64-pc-linux-gnu (64-bit)
R is free software and comes with ABSOLUTELY NO WARRANTY.
You are welcome to redistribute it under certain conditions.
Type 'license()' or 'licence()' for distribution details.
R is a collaborative project with many contributors.
Type 'contributors()' for more information and
'citation()' on how to cite R or R packages in publications.
Type 'demo()' for some demos, 'help()' for on-line help, or
'help.start()' for an HTML browser interface to help.
Type 'q()' to quit R.
> library(OCplus)
Loading required package: akima
> png(filename="/home/ddbj/snapshot/RGM3/R_BC/result/OCplus/plot.fdr1d.result.Rd_%03d_medium.png", width=480, height=480)
> ### Name: plot.fdr1d.result
> ### Title: Plot univariate local false discovery output
> ### Aliases: plot.fdr1d.result
> ### Keywords: hplot aplot
>
> ### ** Examples
>
> example(fdr1d)
fdr1d> # We simulate a small example with 5 percent regulated genes and
fdr1d> # a rather large effect size
fdr1d> set.seed(2000)
fdr1d> xdat = matrix(rnorm(50000), nrow=1000)
fdr1d> xdat[1:25, 1:25] = xdat[1:25, 1:25] - 1
fdr1d> xdat[26:50, 1:25] = xdat[26:50, 1:25] + 1
fdr1d> grp = rep(c("Sample A","Sample B"), c(25,25))
fdr1d> # A default run
fdr1d> res1d = fdr1d(xdat, grp)
Starting permutations...
1
2
3
4
5
6
7
8
9
10
11
12
13
14
15
16
17
18
19
20
21
22
23
24
25
26
27
28
29
30
31
32
33
34
35
36
37
38
39
40
41
42
43
44
45
46
47
48
49
50
51
52
53
54
55
56
57
58
59
60
61
62
63
64
65
66
67
68
69
70
71
72
73
74
75
76
77
78
79
80
81
82
83
84
85
86
87
88
89
90
91
92
93
94
95
96
97
98
99
100
Smoothing f
Smoothing f0
fdr1d> res1d[1:20,]
tstat fdr.local
1 -3.113975 0.343518819
2 -4.868530 0.005661948
3 -4.487968 0.014988680
4 -3.891546 0.067078413
5 -2.323968 0.794599504
6 -2.491116 0.697740540
7 -2.896670 0.460201903
8 -4.254195 0.026995885
9 -3.490224 0.170529676
10 -3.465717 0.180666869
11 -2.642704 0.605687562
12 -1.532776 0.976587492
13 -3.805811 0.082774181
14 -4.138912 0.035586709
15 -4.999346 0.004067203
16 -3.724595 0.098722065
17 -4.853666 0.005843152
18 -2.607185 0.627256785
19 -3.639713 0.123482696
20 -4.881803 0.005500135
fdr1d> # Looking at the results
fdr1d> summary(res1d)
$Statistic
Min. 1st Qu. Median Mean 3rd Qu. Max.
-5.247000 -0.647800 -0.003438 0.015690 0.736200 5.126000
$fdr
fdr
statistic (0,0.05] (0.05,0.1] (0.1,0.2] (0.2,1] (1,Inf]
t<0 8 2 6 480 0
t>=0 11 3 3 487 0
$p0
$p0$Value
[1] 0.8879223
$p0$Estimated
[1] TRUE
fdr1d> plot(res1d)
fdr1d> res1d[res1d$fdr<0.05, ]
tstat fdr.local
2 -4.868530 0.005661948
3 -4.487968 0.014988680
8 -4.254195 0.026995885
14 -4.138912 0.035586709
15 -4.999346 0.004067203
17 -4.853666 0.005843152
20 -4.881803 0.005500135
21 -4.273400 0.025780390
22 -5.247257 0.002150341
23 -4.550108 0.012704082
24 -5.002789 0.004025231
28 4.027545 0.035205867
30 4.274270 0.018487088
32 3.913886 0.047579527
37 4.810875 0.004376113
38 5.125711 0.001711363
39 4.883544 0.003480536
41 4.404355 0.013438726
522 4.017566 0.036292167
fdr1d> # Averaging estimates the global FDR for a set of genes
fdr1d> ndx = abs(res1d$tstat) > 3
fdr1d> mean(res1d$fdr[ndx])
[1] 0.08513626
fdr1d> # Extra information
fdr1d> class(res1d)
[1] "fdr1d.result" "fdr.result" "data.frame"
fdr1d> attr(res1d,"param")
$p0
[1] 0.8879223
$p0.est
[1] TRUE
$fdr
[1] 0.002150341 0.003679441 0.006313912 0.010908497 0.018853690 0.032531021
[7] 0.056260686 0.095823210 0.158862377 0.248254978 0.357555584 0.474579801
[13] 0.598819090 0.730049134 0.852474483 0.928910674 0.961551113 0.977649650
[19] 0.972012961 0.963954436 0.944227575 0.925816181 0.879585482 0.875265249
[25] 0.881280571 0.936401140 0.976673462 1.000000000 0.995431165 0.965258323
[31] 0.930756968 0.906137071 0.900032160 0.843371879 0.769103282 0.673382987
[37] 0.559230197 0.442600785 0.334090209 0.232379026 0.150794383 0.094266063
[43] 0.056810882 0.033284476 0.018990523 0.010603980 0.005822593 0.003159320
[49] 0.001711363
$xbreaks
[1] -5.35530881 -5.13920530 -4.92310180 -4.70699829 -4.49089478 -4.27479127
[7] -4.05868776 -3.84258426 -3.62648075 -3.41037724 -3.19427373 -2.97817022
[13] -2.76206672 -2.54596321 -2.32985970 -2.11375619 -1.89765268 -1.68154918
[19] -1.46544567 -1.24934216 -1.03323865 -0.81713514 -0.60103164 -0.38492813
[25] -0.16882462 0.04727889 0.26338240 0.47948590 0.69558941 0.91169292
[31] 1.12779643 1.34389994 1.56000344 1.77610695 1.99221046 2.20831397
[37] 2.42441748 2.64052098 2.85662449 3.07272800 3.28883151 3.50493502
[43] 3.72103852 3.93714203 4.15324554 4.36934905 4.58545256 4.80155606
[49] 5.01765957 5.23376308
> plot(res1d, grid=TRUE, rug=FALSE)
>
>
>
>
>
> dev.off()
null device
1
>

|
|
 providing
providing  .
.