Supported by Dr. Osamu Ogasawara and  providing providing  . . |
|
Last data update: 2014.03.03 |
Converts matrix to vector based on spot layoutDescriptionThis function converts a matrix based on the spatial layout of spots to a vector. Optionally, a 2D-plot is produced. Usagem2v(M,Ngc,Ngr,Nsc,Nsr,visu=FALSE,color.lim=c(-1,1),xlab="Columns",ylab="Rows",...) Arguments
DetailsThe function ValueA vector Author(s)Matthias E. Futschik (http://itb.biologie.hu-berlin.de/~futschik) See Also
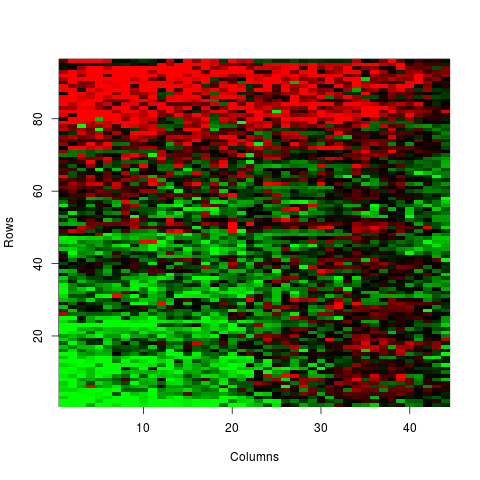
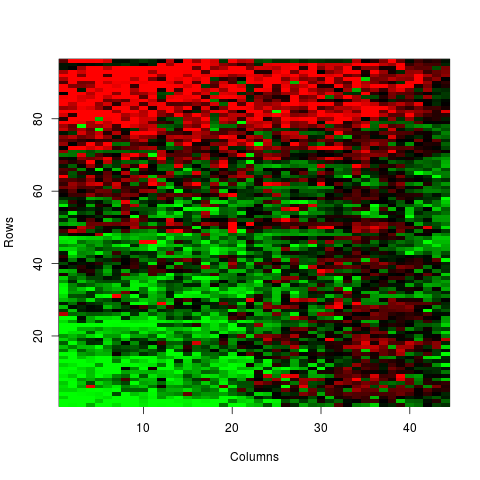
Examples# LOADING DATA NOT-NORMALISED data(sw) # CONVERSION FROM VECTOR TO MATRIX M <- v2m(maM(sw)[,1],Ngc=maNgc(sw),Ngr=maNgr(sw),Nsc=maNsc(sw),Nsr=maNsr(sw),visu=TRUE) # BACK-CONVERSION FROM MATRIX TO VECTOR V <- m2v(M,Ngc=maNgc(sw),Ngr=maNgr(sw),Nsc=maNsc(sw),Nsr=maNsr(sw),visu=TRUE) Results
R version 3.3.1 (2016-06-21) -- "Bug in Your Hair"
Copyright (C) 2016 The R Foundation for Statistical Computing
Platform: x86_64-pc-linux-gnu (64-bit)
R is free software and comes with ABSOLUTELY NO WARRANTY.
You are welcome to redistribute it under certain conditions.
Type 'license()' or 'licence()' for distribution details.
R is a collaborative project with many contributors.
Type 'contributors()' for more information and
'citation()' on how to cite R or R packages in publications.
Type 'demo()' for some demos, 'help()' for on-line help, or
'help.start()' for an HTML browser interface to help.
Type 'q()' to quit R.
> library(OLIN)
Loading required package: locfit
locfit 1.5-9.1 2013-03-22
Loading required package: marray
Loading required package: limma
> png(filename="/home/ddbj/snapshot/RGM3/R_BC/result/OLIN/m2v.Rd_%03d_medium.png", width=480, height=480)
> ### Name: m2v
> ### Title: Converts matrix to vector based on spot layout
> ### Aliases: m2v
> ### Keywords: manip
>
> ### ** Examples
>
>
> # LOADING DATA NOT-NORMALISED
> data(sw)
> # CONVERSION FROM VECTOR TO MATRIX
> M <- v2m(maM(sw)[,1],Ngc=maNgc(sw),Ngr=maNgr(sw),Nsc=maNsc(sw),Nsr=maNsr(sw),visu=TRUE)
>
> # BACK-CONVERSION FROM MATRIX TO VECTOR
> V <- m2v(M,Ngc=maNgc(sw),Ngr=maNgr(sw),Nsc=maNsc(sw),Nsr=maNsr(sw),visu=TRUE)
>
>
>
>
>
> dev.off()
null device
1
>
|