|
Last data update: 2014.03.03
|
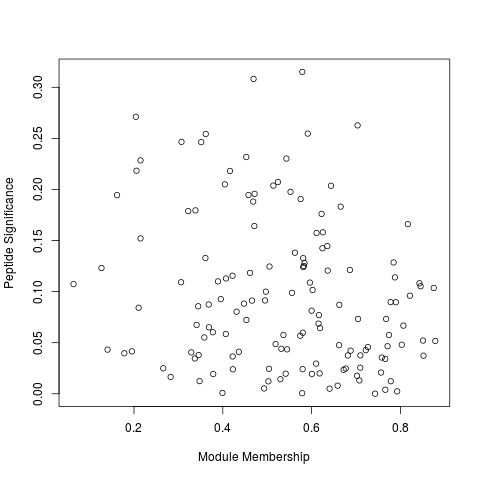
R: Module members vs Peptide Significance
Module members vs Peptide Significance
Description
Plots the module membership (correlation to eigenvector) against the peptide significance (correlation to phenotype) for a given trait and module
Usage
MMvsPS(pnet, pepdat, phenoVec, mod)
Arguments
pnet |
The procona network
|
pepdat |
the peptide data, with rows as samples and columns as peptides
|
phenoVec |
the phenotypic trait, vector
|
mod |
the module of interest
|
Value
returns a list of module memberships and peptide significances.
Author(s)
David L Gibbs
Examples
data(ProCoNA_Data)
#net1 <- buildProconaNetwork("peptide network", peptideData, pow=13)
MMvsPS(net1, peptideData, phenotypes[,5], 1)
Results
R version 3.3.1 (2016-06-21) -- "Bug in Your Hair"
Copyright (C) 2016 The R Foundation for Statistical Computing
Platform: x86_64-pc-linux-gnu (64-bit)
R is free software and comes with ABSOLUTELY NO WARRANTY.
You are welcome to redistribute it under certain conditions.
Type 'license()' or 'licence()' for distribution details.
R is a collaborative project with many contributors.
Type 'contributors()' for more information and
'citation()' on how to cite R or R packages in publications.
Type 'demo()' for some demos, 'help()' for on-line help, or
'help.start()' for an HTML browser interface to help.
Type 'q()' to quit R.
> library(ProCoNA)
Loading required package: WGCNA
Loading required package: dynamicTreeCut
Loading required package: fastcluster
Attaching package: 'fastcluster'
The following object is masked from 'package:stats':
hclust
==========================================================================
*
* Package WGCNA 1.51 loaded.
*
* Important note: It appears that your system supports multi-threading,
* but it is not enabled within WGCNA in R.
* To allow multi-threading within WGCNA with all available cores, use
*
* allowWGCNAThreads()
*
* within R. Use disableWGCNAThreads() to disable threading if necessary.
* Alternatively, set the following environment variable on your system:
*
* ALLOW_WGCNA_THREADS=<number_of_processors>
*
* for example
*
* ALLOW_WGCNA_THREADS=4
*
* To set the environment variable in linux bash shell, type
*
* export ALLOW_WGCNA_THREADS=4
*
* before running R. Other operating systems or shells will
* have a similar command to achieve the same aim.
*
==========================================================================
Attaching package: 'WGCNA'
The following object is masked from 'package:stats':
cor
Loading required package: MSnbase
Loading required package: BiocGenerics
Loading required package: parallel
Attaching package: 'BiocGenerics'
The following objects are masked from 'package:parallel':
clusterApply, clusterApplyLB, clusterCall, clusterEvalQ,
clusterExport, clusterMap, parApply, parCapply, parLapply,
parLapplyLB, parRapply, parSapply, parSapplyLB
The following objects are masked from 'package:stats':
IQR, mad, xtabs
The following objects are masked from 'package:base':
Filter, Find, Map, Position, Reduce, anyDuplicated, append,
as.data.frame, cbind, colnames, do.call, duplicated, eval, evalq,
get, grep, grepl, intersect, is.unsorted, lapply, lengths, mapply,
match, mget, order, paste, pmax, pmax.int, pmin, pmin.int, rank,
rbind, rownames, sapply, setdiff, sort, table, tapply, union,
unique, unsplit
Loading required package: Biobase
Welcome to Bioconductor
Vignettes contain introductory material; view with
'browseVignettes()'. To cite Bioconductor, see
'citation("Biobase")', and for packages 'citation("pkgname")'.
Loading required package: mzR
Loading required package: Rcpp
Loading required package: BiocParallel
Loading required package: ProtGenerics
This is MSnbase version 1.20.7
Read '?MSnbase' and references therein for information
about the package and how to get started.
Attaching package: 'MSnbase'
The following object is masked from 'package:stats':
smooth
The following object is masked from 'package:base':
trimws
Loading required package: flashClust
Attaching package: 'flashClust'
The following object is masked from 'package:fastcluster':
hclust
The following object is masked from 'package:stats':
hclust
Attaching package: 'ProCoNA'
The following object is masked from 'package:ProtGenerics':
peptides
The following object is masked from 'package:Biobase':
samples
> png(filename="/home/ddbj/snapshot/RGM3/R_BC/result/ProCoNA/MMvsPS.Rd_%03d_medium.png", width=480, height=480)
> ### Name: MMvsPS
> ### Title: Module members vs Peptide Significance
> ### Aliases: MMvsPS MMvsPS,proconaNet,matrix,numeric,numeric-method
>
> ### ** Examples
>
> data(ProCoNA_Data)
> #net1 <- buildProconaNetwork("peptide network", peptideData, pow=13)
> MMvsPS(net1, peptideData, phenotypes[,5], 1)
140 peptides in module 1
[[1]]
[1] 0.14053206 0.21046083 0.19512676 0.12682828 0.84287047 0.77797097
[7] 0.84544646 0.80673035 0.79277342 0.87858345 0.82140435 0.75654602
[13] 0.85089781 0.72691490 0.87509465 0.76783423 0.76586750 0.81672274
[19] 0.85208534 0.77833724 0.78765261 0.76601264 0.70970196 0.66242028
[25] 0.80306452 0.77434508 0.68794677 0.75801661 0.61906607 0.78962120
[31] 0.78472667 0.74300043 0.66545153 0.70753850 0.72201817 0.70360720
[37] 0.62529488 0.66193827 0.70231918 0.77111977 0.68627262 0.61031018
[43] 0.59619559 0.65866169 0.55254174 0.63629326 0.57452221 0.61809677
[49] 0.67208965 0.63544609 0.58016904 0.60013028 0.53189468 0.61662045
[55] 0.70990379 0.50275393 0.62444026 0.60074809 0.54486126 0.58263330
[61] 0.50407027 0.49700520 0.57909015 0.51890321 0.52981665 0.61560672
[67] 0.53662060 0.67692503 0.57985085 0.62249868 0.55596796 0.68159883
[73] 0.50520027 0.44782055 0.51399219 0.57535013 0.70466311 0.47158771
[79] 0.46568442 0.58108005 0.49310052 0.36084478 0.64055743 0.45291098
[85] 0.64373134 0.40675276 0.59134532 0.54324195 0.46846619 0.60228112
[91] 0.49519322 0.43104133 0.57879366 0.46130697 0.47111395 0.42259094
[97] 0.42205113 0.36158531 0.20419276 0.58396657 0.35840338 0.43607623
[103] 0.41618774 0.40474574 0.54162491 0.45783252 0.33846588 0.58106003
[109] 0.34104468 0.33704379 0.40729708 0.61118056 0.37763880 0.42222076
[115] 0.06380649 0.39924694 0.38916559 0.56243358 0.34774283 0.36878289
[121] 0.52413953 0.21476051 0.36832781 0.39550786 0.30615275 0.45310864
[127] 0.30697361 0.46929244 0.20563494 0.17828326 0.37846872 0.21466607
[133] 0.32220683 0.35142868 0.34468189 0.34560780 0.32898514 0.16180840
[139] 0.28250655 0.26600698
[[2]]
[1] 4.322923e-02 8.424631e-02 4.153089e-02 1.230254e-01 1.081466e-01
[6] 8.972655e-02 1.052639e-01 6.671107e-02 2.367802e-03 5.172061e-02
[11] 9.600227e-02 2.078354e-02 5.210517e-02 4.565709e-02 1.035774e-01
[16] 7.327028e-02 3.415435e-02 1.660518e-01 3.728287e-02 1.227277e-02
[21] 1.139290e-01 3.946437e-03 2.554295e-02 8.710705e-02 4.800919e-02
[26] 5.756268e-02 4.217262e-02 3.546982e-02 6.429522e-02 8.959064e-02
[31] 1.285206e-01 8.328659e-05 1.831806e-01 1.305131e-02 4.269578e-02
[36] 2.627173e-01 1.580455e-01 4.756029e-02 1.763241e-02 4.653796e-02
[41] 1.211901e-01 2.952572e-02 1.088088e-01 7.915774e-03 1.976287e-01
[46] 1.205586e-01 5.683264e-02 1.992223e-02 2.348652e-02 1.445976e-01
[51] 5.980183e-02 8.133105e-02 4.403556e-02 7.697707e-02 3.755067e-02
[56] 1.217680e-02 1.424041e-01 1.946646e-02 4.348139e-02 1.250646e-01
[61] 2.445118e-02 9.996332e-02 3.150526e-01 4.867194e-02 1.425827e-02
[66] 6.866145e-02 5.756698e-02 2.472134e-02 2.405252e-02 1.760393e-01
[71] 9.879580e-02 3.752052e-02 1.245443e-01 8.822979e-02 2.037829e-01
[76] 1.905975e-01 7.325632e-02 1.956769e-01 9.120662e-02 1.239891e-01
[81] 5.217273e-03 1.328863e-01 5.020140e-03 2.316898e-01 2.036099e-01
[86] 5.851290e-02 2.546911e-01 2.302335e-01 1.881295e-01 1.016296e-01
[91] 9.134052e-02 8.033390e-02 5.463313e-04 1.183327e-01 1.641900e-01
[96] 2.398570e-02 1.155626e-01 2.543223e-01 2.711235e-01 1.279091e-01
[101] 5.512192e-02 4.108621e-02 2.181218e-01 2.049685e-01 1.967993e-02
[106] 1.944794e-01 1.795040e-01 1.327352e-01 6.742574e-02 3.456763e-02
[111] 1.128578e-01 1.573712e-01 6.025355e-02 3.649790e-02 1.072997e-01
[116] 8.235261e-04 1.100052e-01 1.380844e-01 1.237518e-02 6.504321e-02
[121] 2.072498e-01 1.520948e-01 8.737617e-02 9.258602e-02 1.091577e-01
[126] 7.225744e-02 2.465639e-01 3.081872e-01 2.183259e-01 3.963314e-02
[131] 1.929281e-02 2.284424e-01 1.787724e-01 2.463106e-01 8.564392e-02
[136] 3.785286e-02 4.049816e-02 1.944134e-01 1.650577e-02 2.495087e-02
>
>
>
>
>
> dev.off()
null device
1
>
|
|
 providing
providing  .
.