|
Last data update: 2014.03.03
|
R: Plot Pathway-Express result
| plot,peRes,missing-method | R Documentation |
Plot Pathway-Express result
Description
Display a two-way plot using two of the p-values from the Pathway-Express analysis.
Usage
## S4 method for signature 'peRes,missing'
plot(x, y, ..., comb.pv.func = compute.fisher,
adjust.method = "fdr", threshold = 0.05, eps = 1e-06)
## S4 method for signature 'peRes,character'
plot(x, y, ..., comb.pv.func = compute.fisher,
adjust.method = "fdr", threshold = 0.05, eps = 1e-06)
Arguments
x |
an object of type peRes-class
|
y |
vector of two p-values names to be combined using comb.pv.func (default: c("pAcc", "pORA")).
|
... |
Arguments to be passed to methods, such as par.
|
comb.pv.func |
the function to combine the p-values - takes as input a vector of p-values
and returns the combined p-value (default: compute.fisher).
|
adjust.method |
the name of the method to adjust the p-value (see p.adjust)
|
threshold |
corrected p-value threshold
|
eps |
any value smaller than this will be considered as eps (default: 1e-6).
|
Author(s)
Calin Voichita and Sorin Draghici
See Also
pe, summary.peRes, plot,pePathway,missing-method
Examples
# load experiment
load(system.file("extdata/E-GEOD-21942.topTable.RData", package = "ROntoTools"))
fc <- top$logFC[top$adj.P.Val <= .01]
names(fc) <- top$entrez[top$adj.P.Val <= .01]
ref <- top$entrez
# load the set of pathways
kpg <- keggPathwayGraphs("hsa")
kpg <- setEdgeWeights(kpg)
kpg <- setNodeWeights(kpg, defaultWeight = 1)
# perform the pathway analysis (for more accurate results use nboot = 2000)
peRes <- pe(fc, graphs = kpg, ref = ref, nboot = 100, verbose = TRUE)
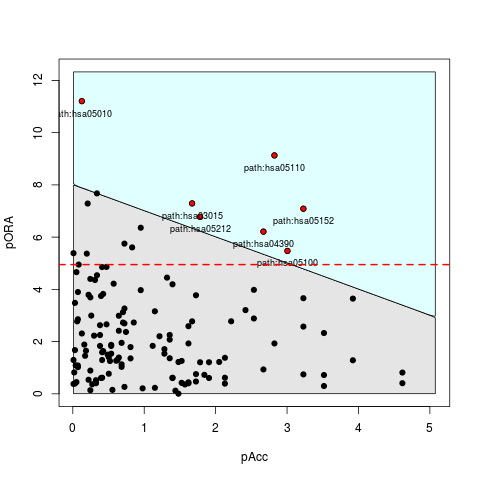
plot(peRes)
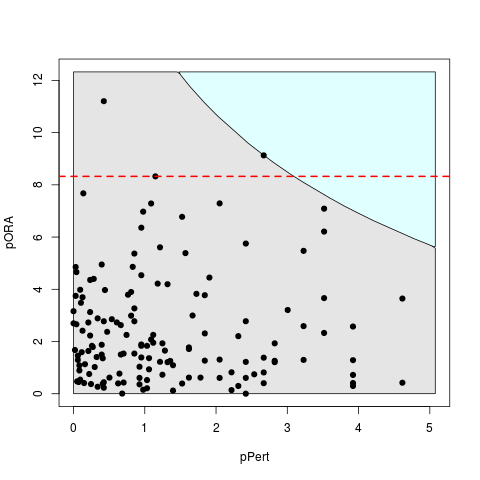
plot(peRes, c("pPert","pORA"), comb.pv.func = compute.normalInv, threshold = .01)
Results
R version 3.3.1 (2016-06-21) -- "Bug in Your Hair"
Copyright (C) 2016 The R Foundation for Statistical Computing
Platform: x86_64-pc-linux-gnu (64-bit)
R is free software and comes with ABSOLUTELY NO WARRANTY.
You are welcome to redistribute it under certain conditions.
Type 'license()' or 'licence()' for distribution details.
R is a collaborative project with many contributors.
Type 'contributors()' for more information and
'citation()' on how to cite R or R packages in publications.
Type 'demo()' for some demos, 'help()' for on-line help, or
'help.start()' for an HTML browser interface to help.
Type 'q()' to quit R.
> library(ROntoTools)
Loading required package: graph
Loading required package: BiocGenerics
Loading required package: parallel
Attaching package: 'BiocGenerics'
The following objects are masked from 'package:parallel':
clusterApply, clusterApplyLB, clusterCall, clusterEvalQ,
clusterExport, clusterMap, parApply, parCapply, parLapply,
parLapplyLB, parRapply, parSapply, parSapplyLB
The following objects are masked from 'package:stats':
IQR, mad, xtabs
The following objects are masked from 'package:base':
Filter, Find, Map, Position, Reduce, anyDuplicated, append,
as.data.frame, cbind, colnames, do.call, duplicated, eval, evalq,
get, grep, grepl, intersect, is.unsorted, lapply, lengths, mapply,
match, mget, order, paste, pmax, pmax.int, pmin, pmin.int, rank,
rbind, rownames, sapply, setdiff, sort, table, tapply, union,
unique, unsplit
Loading required package: boot
Loading required package: KEGGREST
Loading required package: KEGGgraph
Attaching package: 'KEGGgraph'
The following object is masked from 'package:graphics':
plot
Loading required package: Rgraphviz
Loading required package: grid
> png(filename="/home/ddbj/snapshot/RGM3/R_BC/result/ROntoTools/plot.peRes-methods.Rd_%03d_medium.png", width=480, height=480)
> ### Name: plot,peRes,missing-method
> ### Title: Plot Pathway-Express result
> ### Aliases: plot,peRes,character-method plot,peRes,missing-method
>
> ### ** Examples
>
>
> # load experiment
> load(system.file("extdata/E-GEOD-21942.topTable.RData", package = "ROntoTools"))
> fc <- top$logFC[top$adj.P.Val <= .01]
> names(fc) <- top$entrez[top$adj.P.Val <= .01]
> ref <- top$entrez
>
> # load the set of pathways
> kpg <- keggPathwayGraphs("hsa")
Using cached pathway data. Database info:
pathway KEGG Pathway Database
path Release 73.0+/01-03, Jan 15
Kanehisa Laboratories
343,170 entries
Default parameters detected. Using pre-parsed data.
> kpg <- setEdgeWeights(kpg)
> kpg <- setNodeWeights(kpg, defaultWeight = 1)
>
> # perform the pathway analysis (for more accurate results use nboot = 2000)
> peRes <- pe(fc, graphs = kpg, ref = ref, nboot = 100, verbose = TRUE)
Performing pathway analysis...
| | | 0% | | | 1% | |= | 1% | |= | 2% | |== | 3% | |=== | 4% | |=== | 5% | |==== | 5% | |==== | 6% | |===== | 7% | |====== | 8% | |====== | 9% | |======= | 9% | |======= | 10% | |======== | 11% | |======== | 12% | |========= | 13% | |========== | 14% | |========== | 15% | |=========== | 15% | |=========== | 16% | |============ | 17% | |============= | 18% | |============= | 19% | |============== | 19% | |============== | 20% | |=============== | 21% | |================ | 22% | |================ | 23% | |================= | 24% | |================= | 25% | |================== | 26% | |=================== | 27% | |=================== | 28% | |==================== | 28% | |==================== | 29% | |===================== | 30% | |====================== | 31% | |====================== | 32% | |======================= | 32% | |======================= | 33% | |======================= | 34% | |======================== | 34% | |======================== | 35% | |========================= | 36% | |========================== | 37% | |========================== | 38% | |=========================== | 38% | |=========================== | 39% | |============================ | 40% | |============================= | 41% | |============================= | 42% | |============================== | 42% | |============================== | 43% | |=============================== | 44% | |=============================== | 45% | |================================ | 46% | |================================= | 47% | |================================= | 48% | |================================== | 48% | |================================== | 49% | |=================================== | 50% | |==================================== | 51% | |==================================== | 52% | |===================================== | 52% | |===================================== | 53% | |====================================== | 54% | |======================================= | 55% | |======================================= | 56% | |======================================== | 57% | |======================================== | 58% | |========================================= | 58% | |========================================= | 59% | |========================================== | 60% | |=========================================== | 61% | |=========================================== | 62% | |============================================ | 62% | |============================================ | 63% | |============================================= | 64% | |============================================== | 65% | |============================================== | 66% | |=============================================== | 66% | |=============================================== | 67% | |=============================================== | 68% | |================================================ | 68% | |================================================ | 69% | |================================================= | 70% | |================================================== | 71% | |================================================== | 72% | |=================================================== | 72% | |=================================================== | 73% | |==================================================== | 74% | |===================================================== | 75% | |===================================================== | 76% | |====================================================== | 77% | |====================================================== | 78% | |======================================================= | 79% | |======================================================== | 80% | |======================================================== | 81% | |========================================================= | 81% | |========================================================= | 82% | |========================================================== | 83% | |=========================================================== | 84% | |=========================================================== | 85% | |============================================================ | 85% | |============================================================ | 86% | |============================================================= | 87% | |============================================================== | 88% | |============================================================== | 89% | |=============================================================== | 90% | |=============================================================== | 91% | |================================================================ | 91% | |================================================================ | 92% | |================================================================= | 93% | |================================================================== | 94% | |================================================================== | 95% | |=================================================================== | 95% | |=================================================================== | 96% | |==================================================================== | 97% | |===================================================================== | 98% | |===================================================================== | 99% | |======================================================================| 99% | |======================================================================| 100%Analysis completed in 4.852925 secs.
>
> plot(peRes)
>
> plot(peRes, c("pPert","pORA"), comb.pv.func = compute.normalInv, threshold = .01)
Warning message:
In max(st[i, "pComb"]) : no non-missing arguments to max; returning -Inf
>
>
>
>
>
>
> dev.off()
null device
1
>
|
|
 providing
providing  .
.