Supported by Dr. Osamu Ogasawara and  providing providing  . . |
|
Last data update: 2014.03.03 |
Extract Survival Information From RTCGA.clinical DatasetsDescriptionExtracts survival information from clicnial datasets from TCGA project. UsagesurvivalTCGA(..., extract.cols = NULL, extract.names = FALSE, barcode.name = "patient.bcr_patient_barcode", event.name = "patient.vital_status", days.to.followup.name = "patient.days_to_last_followup", days.to.death.name = "patient.days_to_death") Arguments
ValueA data.frame containing information about times and censoring for specific IssuesIf you have any problems, issues or think that something is missing or is not clear please post an issue on https://github.com/RTCGA/RTCGA/issues. NoteInput data.frames should contain columns Author(s)Marcin Kosinski, m.p.kosinski@gmail.com Marcin Kosinski, m.p.kosinski@gmail.com See AlsoRTCGA website http://rtcga.github.io/RTCGA/Visualizations.html. Other RTCGA: Examples
## Extracting Survival Data
library(RTCGA.clinical)
survivalTCGA(BRCA.clinical, OV.clinical, extract.cols = "admin.disease_code") -> BRCAOV.survInfo
# first munge data, then extract survival info
library(dplyr)
BRCA.clinical %>%
filter(patient.drugs.drug.therapy_types.therapy_type %in%
c("chemotherapy", "hormone therapy")) %>%
rename(therapy = patient.drugs.drug.therapy_types.therapy_type) %>%
survivalTCGA(extract.cols = c("therapy")) -> BRCA.survInfo.chemo
# first extract survival info, then munge data
survivalTCGA(BRCA.clinical,
extract.cols = c("patient.drugs.drug.therapy_types.therapy_type")) %>%
filter(patient.drugs.drug.therapy_types.therapy_type %in%
c("chemotherapy", "hormone therapy")) %>%
rename(therapy = patient.drugs.drug.therapy_types.therapy_type) -> BRCA.survInfo.chemo
## Kaplan-Meier Survival Curves
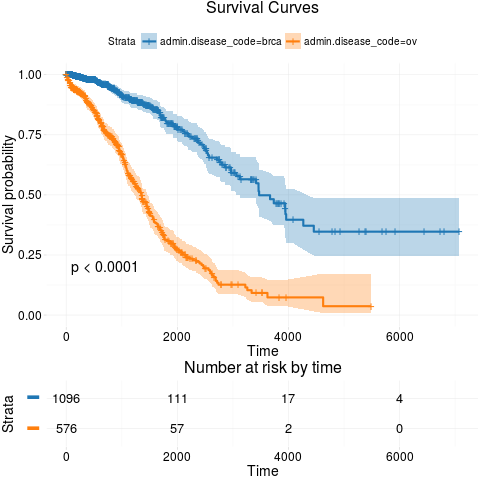
kmTCGA(BRCAOV.survInfo, explanatory.names = "admin.disease_code", pval = TRUE)
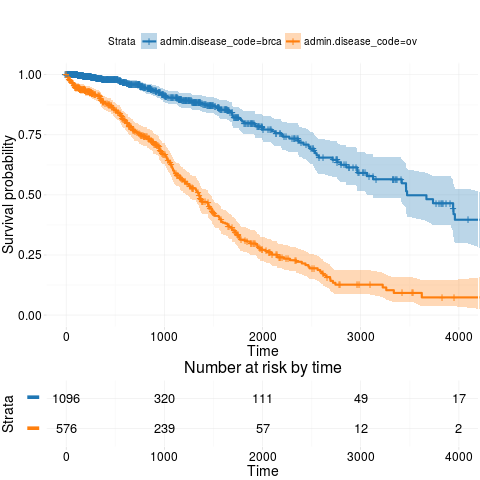
kmTCGA(BRCAOV.survInfo, explanatory.names = "admin.disease_code", main = "",
xlim = c(0,4000))
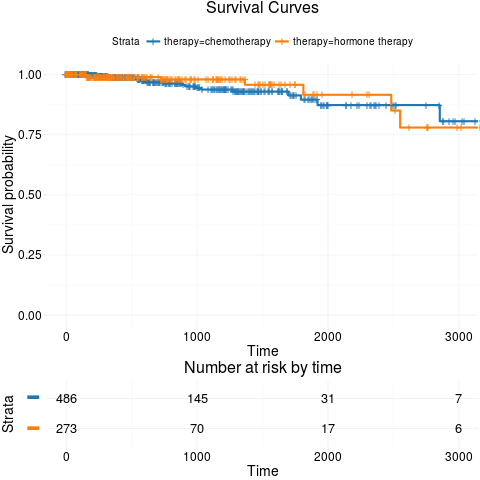
kmTCGA(BRCA.survInfo.chemo, explanatory.names = "therapy", xlim = c(0, 3000), conf.int = FALSE)
Results
R version 3.3.1 (2016-06-21) -- "Bug in Your Hair"
Copyright (C) 2016 The R Foundation for Statistical Computing
Platform: x86_64-pc-linux-gnu (64-bit)
R is free software and comes with ABSOLUTELY NO WARRANTY.
You are welcome to redistribute it under certain conditions.
Type 'license()' or 'licence()' for distribution details.
R is a collaborative project with many contributors.
Type 'contributors()' for more information and
'citation()' on how to cite R or R packages in publications.
Type 'demo()' for some demos, 'help()' for on-line help, or
'help.start()' for an HTML browser interface to help.
Type 'q()' to quit R.
> library(RTCGA)
Welcome to the RTCGA (version: 1.2.2).
> png(filename="/home/ddbj/snapshot/RGM3/R_BC/result/RTCGA/survivalTCGA.Rd_%03d_medium.png", width=480, height=480)
> ### Name: survivalTCGA
> ### Title: Extract Survival Information From RTCGA.clinical Datasets
> ### Aliases: survivalTCGA
>
> ### ** Examples
>
>
> ## Extracting Survival Data
> library(RTCGA.clinical)
> survivalTCGA(BRCA.clinical, OV.clinical, extract.cols = "admin.disease_code") -> BRCAOV.survInfo
>
> # first munge data, then extract survival info
> library(dplyr)
Attaching package: 'dplyr'
The following objects are masked from 'package:stats':
filter, lag
The following objects are masked from 'package:base':
intersect, setdiff, setequal, union
> BRCA.clinical %>%
+ filter(patient.drugs.drug.therapy_types.therapy_type %in%
+ c("chemotherapy", "hormone therapy")) %>%
+ rename(therapy = patient.drugs.drug.therapy_types.therapy_type) %>%
+ survivalTCGA(extract.cols = c("therapy")) -> BRCA.survInfo.chemo
>
> # first extract survival info, then munge data
> survivalTCGA(BRCA.clinical,
+ extract.cols = c("patient.drugs.drug.therapy_types.therapy_type")) %>%
+ filter(patient.drugs.drug.therapy_types.therapy_type %in%
+ c("chemotherapy", "hormone therapy")) %>%
+ rename(therapy = patient.drugs.drug.therapy_types.therapy_type) -> BRCA.survInfo.chemo
>
> ## Kaplan-Meier Survival Curves
> kmTCGA(BRCAOV.survInfo, explanatory.names = "admin.disease_code", pval = TRUE)
>
> kmTCGA(BRCAOV.survInfo, explanatory.names = "admin.disease_code", main = "",
+ xlim = c(0,4000))
>
> kmTCGA(BRCA.survInfo.chemo, explanatory.names = "therapy", xlim = c(0, 3000), conf.int = FALSE)
>
>
>
>
>
>
> dev.off()
null device
1
>
|