Supported by Dr. Osamu Ogasawara and  providing providing  . . |
|
Last data update: 2014.03.03 |
CNV.genomeplotDescriptionCreate CNV plot for the whole genome or chromosomes. Usage
CNV.genomeplot(object, ...)
## S4 method for signature 'CNV.analysis'
CNV.genomeplot(object, chr = "all", chrX = TRUE,
chrY = TRUE, centromere = TRUE, detail = TRUE, main = NULL,
ylim = c(-1.25, 1.25), set_par = TRUE, cols = c("red", "red",
"lightgrey", "green", "green"))
## S4 method for signature 'CNV.analysis,ANY'
plot(x, y = NULL, ...)
Arguments
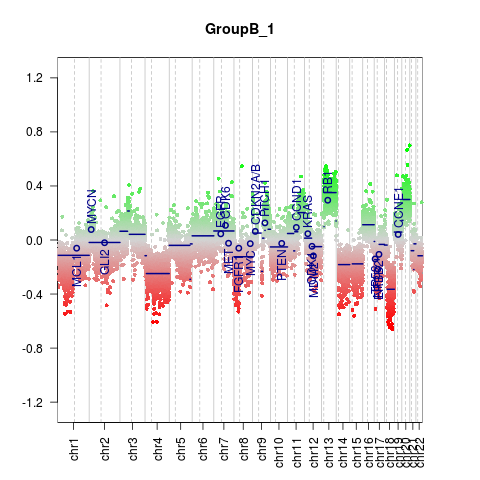
DetailsThis method provides the functionality for generating CNV plots for the whole genome or defined chromosomes. Bins are shown as dots, segments are shown as lines. See parameters for more information. Value
Author(s)Volker Hovestadt conumee@hovestadt.bio Examples
# prepare
library(minfiData)
data(MsetEx)
d <- CNV.load(MsetEx)
data(detail_regions)
anno <- CNV.create_anno(detail_regions = detail_regions)
# create/modify object
x <- CNV.segment(CNV.detail(CNV.bin(CNV.fit(query = d['GroupB_1'],
ref = d[c('GroupA_1', 'GroupA_2', 'GroupA_3')], anno))))
# output plots
CNV.genomeplot(x)
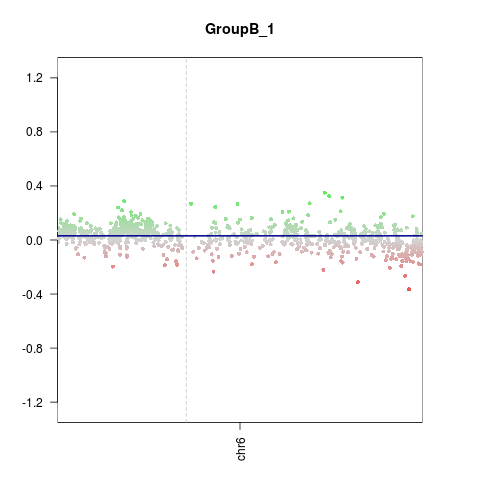
CNV.genomeplot(x, chr = 'chr6')
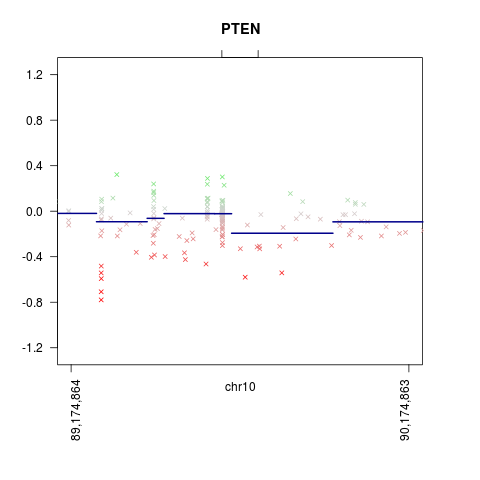
CNV.detailplot(x, name = 'PTEN')
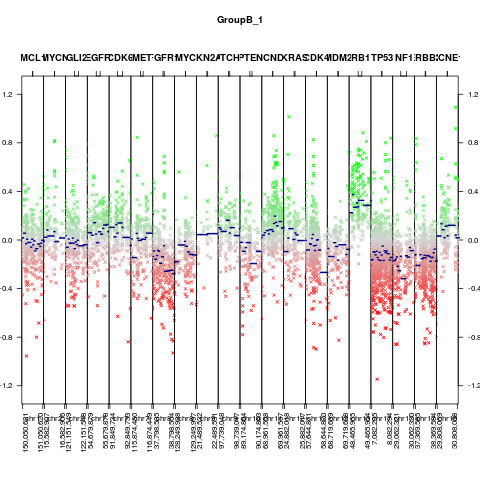
CNV.detailplot_wrap(x)
# output text files
CNV.write(x, what = 'segments')
CNV.write(x, what = 'detail')
CNV.write(x, what = 'bins')
CNV.write(x, what = 'probes')
Results
R version 3.3.1 (2016-06-21) -- "Bug in Your Hair"
Copyright (C) 2016 The R Foundation for Statistical Computing
Platform: x86_64-pc-linux-gnu (64-bit)
R is free software and comes with ABSOLUTELY NO WARRANTY.
You are welcome to redistribute it under certain conditions.
Type 'license()' or 'licence()' for distribution details.
R is a collaborative project with many contributors.
Type 'contributors()' for more information and
'citation()' on how to cite R or R packages in publications.
Type 'demo()' for some demos, 'help()' for on-line help, or
'help.start()' for an HTML browser interface to help.
Type 'q()' to quit R.
> library(conumee)
Loading required package: minfi
Loading required package: BiocGenerics
Loading required package: parallel
Attaching package: 'BiocGenerics'
The following objects are masked from 'package:parallel':
clusterApply, clusterApplyLB, clusterCall, clusterEvalQ,
clusterExport, clusterMap, parApply, parCapply, parLapply,
parLapplyLB, parRapply, parSapply, parSapplyLB
The following objects are masked from 'package:stats':
IQR, mad, xtabs
The following objects are masked from 'package:base':
Filter, Find, Map, Position, Reduce, anyDuplicated, append,
as.data.frame, cbind, colnames, do.call, duplicated, eval, evalq,
get, grep, grepl, intersect, is.unsorted, lapply, lengths, mapply,
match, mget, order, paste, pmax, pmax.int, pmin, pmin.int, rank,
rbind, rownames, sapply, setdiff, sort, table, tapply, union,
unique, unsplit
Loading required package: Biobase
Welcome to Bioconductor
Vignettes contain introductory material; view with
'browseVignettes()'. To cite Bioconductor, see
'citation("Biobase")', and for packages 'citation("pkgname")'.
Loading required package: lattice
Loading required package: GenomicRanges
Loading required package: S4Vectors
Loading required package: stats4
Attaching package: 'S4Vectors'
The following objects are masked from 'package:base':
colMeans, colSums, expand.grid, rowMeans, rowSums
Loading required package: IRanges
Loading required package: GenomeInfoDb
Loading required package: SummarizedExperiment
Loading required package: Biostrings
Loading required package: XVector
Loading required package: bumphunter
Loading required package: foreach
Loading required package: iterators
Loading required package: locfit
locfit 1.5-9.1 2013-03-22
Setting options('download.file.method.GEOquery'='auto')
Setting options('GEOquery.inmemory.gpl'=FALSE)
Loading required package: IlluminaHumanMethylation450kmanifest
Loading required package: IlluminaHumanMethylation450kanno.ilmn12.hg19
> png(filename="/home/ddbj/snapshot/RGM3/R_BC/result/conumee/CNV.genomeplot.Rd_%03d_medium.png", width=480, height=480)
> ### Name: CNV.genomeplot
> ### Title: CNV.genomeplot
> ### Aliases: CNV.genomeplot CNV.genomeplot,CNV.analysis-method
> ### plot,CNV.analysis,ANY-method
>
> ### ** Examples
>
> # prepare
> library(minfiData)
> data(MsetEx)
> d <- CNV.load(MsetEx)
> data(detail_regions)
> anno <- CNV.create_anno(detail_regions = detail_regions)
using genome annotations from UCSC
getting 450k annotations
- 470870 probes used
importing regions for detailed analysis
creating bins
- 53918 bins created
merging bins
- 15833 bins remaining
>
> # create/modify object
> x <- CNV.segment(CNV.detail(CNV.bin(CNV.fit(query = d['GroupB_1'],
+ ref = d[c('GroupA_1', 'GroupA_2', 'GroupA_3')], anno))))
>
> # output plots
> CNV.genomeplot(x)
> CNV.genomeplot(x, chr = 'chr6')
> CNV.detailplot(x, name = 'PTEN')
> CNV.detailplot_wrap(x)
>
> # output text files
> CNV.write(x, what = 'segments')
ID chrom loc.start loc.end num.mark bstat pval seg.mean
1 GroupB_1 chr1 635684 249195311 1593 NA NA -0.122
2 GroupB_1 chr10 105000 135462374 840 NA NA -0.058
3 GroupB_1 chr11 130000 48550000 319 7.851693 4.823678e-13 0.042
4 GroupB_1 chr11 48975000 56675000 15 12.294071 1.649339e-32 -0.091
5 GroupB_1 chr11 56975000 81250000 268 29.745363 9.783071e-192 0.057
6 GroupB_1 chr11 82500000 134873258 312 NA NA 0.199
7 GroupB_1 chr12 172870 54975000 351 11.507897 2.178154e-28 0.011
8 GroupB_1 chr12 55175000 56075000 5 8.018864 1.507192e-13 -0.178
9 GroupB_1 chr12 56125000 133770948 492 NA NA -0.064
10 GroupB_1 chr13 19110000 19525000 3 5.415424 4.913394e-06 0.119
11 GroupB_1 chr13 19725000 114025000 415 9.565066 1.824189e-19 0.348
12 GroupB_1 chr13 114075000 115079939 13 NA NA 0.151
13 GroupB_1 chr14 19325000 107119770 528 NA NA -0.197
14 GroupB_1 chr15 20225000 102410696 571 NA NA -0.189
15 GroupB_1 chr16 105000 90222377 624 NA NA 0.114
16 GroupB_1 chr17 25000 3875000 59 16.221715 8.901516e-57 -0.015
17 GroupB_1 chr17 3925000 38850000 317 7.820936 6.065279e-13 -0.160
18 GroupB_1 chr17 38925000 39400000 8 16.150879 3.079978e-56 -0.345
19 GroupB_1 chr17 39475000 81122605 430 NA NA -0.053
20 GroupB_1 chr18 80000 21125000 81 21.391647 3.413057e-99 -0.031
21 GroupB_1 chr18 21375000 77983624 162 NA NA -0.364
22 GroupB_1 chr19 180000 59084492 711 NA NA 0.050
23 GroupB_1 chr2 130000 243026238 1282 NA NA -0.027
24 GroupB_1 chr20 155000 8725000 65 13.487427 3.902028e-39 0.154
25 GroupB_1 chr20 9475000 62907760 294 NA NA 0.297
26 GroupB_1 chr21 10848948 31025000 27 5.747398 2.031374e-07 -0.073
27 GroupB_1 chr21 31525000 32125000 5 8.799096 1.101514e-16 -0.269
28 GroupB_1 chr21 32575000 48084948 125 NA NA -0.032
29 GroupB_1 chr22 16373925 51222283 300 NA NA -0.122
30 GroupB_1 chr3 155000 60150000 362 18.594996 1.060849e-74 0.056
31 GroupB_1 chr3 60850000 73675000 64 18.946425 1.813659e-77 0.215
32 GroupB_1 chr3 74175000 197831215 552 NA NA 0.035
33 GroupB_1 chr4 55000 10475000 136 14.363704 2.842430e-44 -0.125
34 GroupB_1 chr4 10925000 190972138 606 NA NA -0.253
35 GroupB_1 chr5 55000 162925000 709 15.310711 2.214461e-50 -0.046
36 GroupB_1 chr5 163475000 167325000 6 10.853541 1.884955e-25 -0.284
37 GroupB_1 chr5 167575000 180777630 134 NA NA -0.038
38 GroupB_1 chr6 155000 170952534 988 NA NA 0.024
39 GroupB_1 chr7 55000 159039332 952 NA NA 0.064
40 GroupB_1 chr8 105000 146277011 726 NA NA -0.133
41 GroupB_1 chr9 130000 8300000 20 10.897739 7.644010e-26 0.204
42 GroupB_1 chr9 10800000 68326090 60 16.752606 3.851767e-61 0.044
43 GroupB_1 chr9 71067734 79450000 17 10.918159 2.924101e-26 -0.235
44 GroupB_1 chr9 79625000 90250000 19 14.633854 7.577804e-47 0.014
45 GroupB_1 chr9 90475000 97450000 30 12.522876 2.179260e-34 0.225
46 GroupB_1 chr9 97650000 99925000 14 6.558129 2.436444e-09 0.074
47 GroupB_1 chr9 100150000 123625000 60 15.832035 3.243051e-54 0.217
48 GroupB_1 chr9 123675000 141076716 163 NA NA 0.083
seg.median
1 -0.114
2 -0.051
3 0.048
4 -0.081
5 0.061
6 0.203
7 0.014
8 -0.198
9 -0.049
10 0.099
11 0.351
12 0.141
13 -0.182
14 -0.177
15 0.113
16 -0.011
17 -0.145
18 -0.354
19 -0.033
20 -0.038
21 -0.365
22 0.059
23 -0.018
24 0.146
25 0.300
26 -0.079
27 -0.219
28 -0.028
29 -0.117
30 0.065
31 0.215
32 0.042
33 -0.116
34 -0.249
35 -0.040
36 -0.291
37 -0.029
38 0.031
39 0.067
40 -0.124
41 0.183
42 0.051
43 -0.232
44 0.014
45 0.230
46 0.071
47 0.227
48 0.079
> CNV.write(x, what = 'detail')
chr start end name sample probes.Var1 probes.Freq value
1 chr1 150547028 150554214 MCL1 GroupB_1 MCL1 18 -0.061
2 chr2 16078684 16087129 MYCN GroupB_1 MYCN 25 0.077
3 chr2 121552868 121750229 GLI2 GroupB_1 GLI2 86 -0.020
4 chr7 55084726 55275031 EGFR GroupB_1 EGFR 57 0.046
5 chr7 92234236 92465231 CDK6 GroupB_1 CDK6 54 0.108
6 chr7 116310460 116438440 MET GroupB_1 MET 24 -0.024
7 chr8 38268657 38328352 FGFR1 GroupB_1 FGFR1 36 -0.061
8 chr8 128746316 128753680 MYC GroupB_1 MYC 39 -0.027
9 chr9 21967752 22011312 CDKN2A/B GroupB_1 CDKN2A/B 14 0.064
10 chr9 98205265 98272831 PTCH1 GroupB_1 PTCH1 24 0.127
11 chr10 89621196 89728532 PTEN GroupB_1 PTEN 65 -0.025
12 chr11 69453874 69469242 CCND1 GroupB_1 CCND1 70 0.093
13 chr12 25358181 25405854 KRAS GroupB_1 KRAS 33 0.047
14 chr12 58141511 58148230 CDK4 GroupB_1 CDK4 24 -0.047
15 chr12 69199953 69239324 MDM2 GroupB_1 MDM2 15 -0.118
16 chr13 48875884 49056026 RB1 GroupB_1 RB1 61 0.294
17 chr17 7571721 7592868 TP53 GroupB_1 TP53 39 -0.140
18 chr17 29419946 29704695 NF1 GroupB_1 NF1 42 -0.216
19 chr17 37854255 37884915 ERBB2 GroupB_1 ERBB2 26 -0.106
20 chr19 30300902 30315215 CCNE1 GroupB_1 CCNE1 16 0.041
> CNV.write(x, what = 'bins')
Chromosome Start End Feature GroupB_1
1 chr1 521369 750000 chr1-0001 -0.103
2 chr1 750001 800000 chr1-0002 -0.125
3 chr1 800001 850000 chr1-0003 -0.149
4 chr1 850001 900000 chr1-0004 -0.079
5 chr1 900001 950000 chr1-0005 -0.069
6 chr1 950001 1000000 chr1-0006 -0.069
7 chr1 1000001 1050000 chr1-0007 -0.095
8 chr1 1050001 1100000 chr1-0008 -0.088
9 chr1 1100001 1150000 chr1-0009 -0.096
10 chr1 1150001 1200000 chr1-0010 -0.060
11 chr1 1200001 1250000 chr1-0011 -0.085
12 chr1 1250001 1300000 chr1-0012 -0.075
13 chr1 1300001 1350000 chr1-0013 -0.071
14 chr1 1350001 1400000 chr1-0014 -0.116
15 chr1 1400001 1450000 chr1-0015 -0.078
16 chr1 1450001 1500000 chr1-0016 -0.069
17 chr1 1500001 1550000 chr1-0017 -0.103
18 chr1 1550001 1600000 chr1-0018 -0.056
19 chr1 1600001 1650000 chr1-0019 -0.154
20 chr1 1650001 1700000 chr1-0020 -0.105
21 chr1 1700001 1800000 chr1-0021 -0.129
22 chr1 1800001 1850000 chr1-0022 -0.095
23 chr1 1850001 1900000 chr1-0023 -0.114
24 chr1 1900001 1950000 chr1-0024 -0.107
25 chr1 1950001 2000000 chr1-0025 -0.060
26 chr1 2000001 2050000 chr1-0026 -0.077
27 chr1 2050001 2100000 chr1-0027 -0.113
28 chr1 2100001 2150000 chr1-0028 -0.084
29 chr1 2150001 2200000 chr1-0029 -0.106
30 chr1 2200001 2250000 chr1-0030 -0.095
31 chr1 2250001 2300000 chr1-0031 -0.065
32 chr1 2300001 2350000 chr1-0032 -0.080
33 chr1 2350001 2400000 chr1-0033 -0.040
34 chr1 2400001 2450000 chr1-0034 -0.049
35 chr1 2450001 2500000 chr1-0035 -0.084
36 chr1 2500001 2550000 chr1-0036 -0.116
37 chr1 2550001 2634220 chr1-0037 -0.066
38 chr1 2684221 2750000 chr1-0038 -0.116
39 chr1 2750001 2800000 chr1-0039 -0.093
40 chr1 2800001 2850000 chr1-0040 -0.120
41 chr1 2850001 2900000 chr1-0041 -0.117
42 chr1 2900001 2950000 chr1-0042 -0.078
43 chr1 2950001 3000000 chr1-0043 -0.097
44 chr1 3000001 3050000 chr1-0044 -0.101
45 chr1 3050001 3100000 chr1-0045 -0.077
46 chr1 3100001 3150000 chr1-0046 -0.131
47 chr1 3150001 3200000 chr1-0047 -0.088
48 chr1 3200001 3250000 chr1-0048 -0.123
49 chr1 3250001 3300000 chr1-0049 -0.079
50 chr1 3300001 3350000 chr1-0050 -0.114
51 chr1 3350001 3400000 chr1-0051 -0.093
52 chr1 3400001 3450000 chr1-0052 -0.053
53 chr1 3450001 3500000 chr1-0053 -0.083
54 chr1 3500001 3550000 chr1-0054 -0.095
55 chr1 3550001 3600000 chr1-0055 -0.073
56 chr1 3600001 3650000 chr1-0056 -0.061
57 chr1 3650001 3700000 chr1-0057 -0.060
58 chr1 3700001 3750000 chr1-0058 -0.134
59 chr1 3750001 3800000 chr1-0059 -0.105
60 chr1 3800001 3845268 chr1-0060 -0.116
61 chr1 3995269 4150000 chr1-0061 -0.100
62 chr1 4150001 4200000 chr1-0062 -0.102
63 chr1 4200001 4450000 chr1-0063 -0.220
64 chr1 4450001 4500000 chr1-0064 -0.349
65 chr1 4500001 4700000 chr1-0065 -0.104
66 chr1 4700001 4750000 chr1-0066 -0.139
67 chr1 4750001 4800000 chr1-0067 -0.115
68 chr1 4800001 4850000 chr1-0068 -0.206
69 chr1 4850001 5550000 chr1-0069 -0.232
70 chr1 5550001 5750000 chr1-0070 -0.142
71 chr1 5750001 5800000 chr1-0071 -0.122
72 chr1 5800001 5900000 chr1-0072 -0.225
73 chr1 5900001 5950000 chr1-0073 -0.096
74 chr1 5950001 6050000 chr1-0074 -0.121
75 chr1 6050001 6100000 chr1-0075 -0.121
76 chr1 6100001 6150000 chr1-0076 -0.096
77 chr1 6150001 6200000 chr1-0077 -0.060
78 chr1 6200001 6250000 chr1-0078 -0.152
79 chr1 6250001 6300000 chr1-0079 -0.120
80 chr1 6300001 6350000 chr1-0080 -0.070
81 chr1 6350001 6400000 chr1-0081 -0.130
82 chr1 6400001 6450000 chr1-0082 -0.170
83 chr1 6450001 6500000 chr1-0083 -0.090
84 chr1 6500001 6550000 chr1-0084 -0.100
85 chr1 6550001 6600000 chr1-0085 -0.116
86 chr1 6600001 6650000 chr1-0086 -0.057
87 chr1 6650001 6700000 chr1-0087 -0.073
88 chr1 6700001 6800000 chr1-0088 -0.206
89 chr1 6800001 6850000 chr1-0089 -0.257
90 chr1 6850001 7000000 chr1-0090 -0.235
91 chr1 7000001 7100000 chr1-0091 -0.158
92 chr1 7100001 7150000 chr1-0092 -0.181
93 chr1 7150001 7300000 chr1-0093 -0.245
94 chr1 7300001 7450000 chr1-0094 -0.187
95 chr1 7450001 7550000 chr1-0095 -0.099
96 chr1 7550001 7700000 chr1-0096 -0.130
97 chr1 7700001 7750000 chr1-0097 -0.095
98 chr1 7750001 7800000 chr1-0098 -0.132
99 chr1 7800001 7850000 chr1-0099 -0.165
100 chr1 7850001 8000000 chr1-0100 -0.242
101 chr1 8000001 8050000 chr1-0101 -0.158
102 chr1 8050001 8100000 chr1-0102 -0.212
103 chr1 8100001 8250000 chr1-0103 -0.336
104 chr1 8250001 8350000 chr1-0104 -0.149
105 chr1 8350001 8400000 chr1-0105 -0.120
106 chr1 8400001 8450000 chr1-0106 -0.107
107 chr1 8450001 8500000 chr1-0107 -0.224
108 chr1 8500001 8700000 chr1-0108 -0.163
109 chr1 8700001 8850000 chr1-0109 -0.233
110 chr1 8850001 8900000 chr1-0110 -0.129
111 chr1 8900001 8950000 chr1-0111 -0.136
112 chr1 8950001 9050000 chr1-0112 -0.196
113 chr1 9050001 9100000 chr1-0113 -0.175
114 chr1 9100001 9150000 chr1-0114 -0.147
115 chr1 9150001 9200000 chr1-0115 -0.221
116 chr1 9200001 9250000 chr1-0116 -0.116
117 chr1 9250001 9300000 chr1-0117 -0.133
118 chr1 9300001 9350000 chr1-0118 -0.087
119 chr1 9350001 9400000 chr1-0119 -0.149
120 chr1 9400001 9450000 chr1-0120 -0.112
121 chr1 9450001 9550000 chr1-0121 -0.224
122 chr1 9550001 9600000 chr1-0122 -0.120
123 chr1 9600001 9650000 chr1-0123 -0.221
124 chr1 9650001 9700000 chr1-0124 -0.110
125 chr1 9700001 9750000 chr1-0125 -0.193
126 chr1 9750001 9800000 chr1-0126 -0.115
127 chr1 9800001 9900000 chr1-0127 -0.157
128 chr1 9900001 10000000 chr1-0128 -0.092
129 chr1 10000001 10050000 chr1-0129 -0.079
130 chr1 10050001 10100000 chr1-0130 -0.179
131 chr1 10100001 10350000 chr1-0131 -0.131
132 chr1 10350001 10450000 chr1-0132 -0.161
133 chr1 10450001 10500000 chr1-0133 -0.125
134 chr1 10500001 10550000 chr1-0134 -0.143
135 chr1 10550001 10650000 chr1-0135 -0.228
136 chr1 10650001 10700000 chr1-0136 -0.119
137 chr1 10700001 10750000 chr1-0137 -0.089
138 chr1 10750001 10800000 chr1-0138 -0.120
139 chr1 10800001 10850000 chr1-0139 -0.138
140 chr1 10850001 10900000 chr1-0140 -0.133
141 chr1 10900001 11000000 chr1-0141 -0.154
142 chr1 11000001 11100000 chr1-0142 -0.117
143 chr1 11100001 11150000 chr1-0143 -0.075
144 chr1 11150001 11300000 chr1-0144 -0.186
145 chr1 11300001 11350000 chr1-0145 -0.132
146 chr1 11350001 11500000 chr1-0146 -0.179
147 chr1 11500001 11550000 chr1-0147 -0.099
148 chr1 11550001 11700000 chr1-0148 -0.096
149 chr1 11700001 11750000 chr1-0149 -0.114
150 chr1 11750001 11800000 chr1-0150 -0.151
151 chr1 11800001 11850000 chr1-0151 -0.042
152 chr1 11850001 11900000 chr1-0152 -0.113
153 chr1 11900001 11950000 chr1-0153 -0.070
154 chr1 11950001 12000000 chr1-0154 -0.096
155 chr1 12000001 12050000 chr1-0155 -0.106
156 chr1 12050001 12100000 chr1-0156 -0.124
157 chr1 12100001 12150000 chr1-0157 -0.271
158 chr1 12150001 12200000 chr1-0158 -0.171
159 chr1 12200001 12250000 chr1-0159 -0.173
160 chr1 12250001 12300000 chr1-0160 -0.106
161 chr1 12300001 12500000 chr1-0161 -0.295
162 chr1 12500001 12550000 chr1-0162 -0.277
163 chr1 12550001 12600000 chr1-0163 -0.089
164 chr1 12600001 12650000 chr1-0164 -0.162
165 chr1 12650001 12700000 chr1-0165 -0.153
166 chr1 12700001 12800000 chr1-0166 -0.229
167 chr1 12800001 12900000 chr1-0167 -0.160
168 chr1 12900001 13052998 chr1-0168 -0.240
169 chr1 13607163 13900000 chr1-0169 -0.140
170 chr1 13900001 14000000 chr1-0170 -0.127
171 chr1 14000001 14050000 chr1-0171 -0.264
172 chr1 14050001 14100000 chr1-0172 -0.167
173 chr1 14100001 14150000 chr1-0173 -0.262
174 chr1 14150001 14550000 chr1-0174 -0.275
175 chr1 14550001 14900000 chr1-0175 -0.232
176 chr1 14900001 14950000 chr1-0176 -0.012
177 chr1 14950001 15150000 chr1-0177 -0.229
178 chr1 15150001 15250000 chr1-0178 -0.232
179 chr1 15250001 15300000 chr1-0179 -0.202
180 chr1 15300001 15450000 chr1-0180 -0.136
181 chr1 15450001 15500000 chr1-0181 -0.203
182 chr1 15500001 15550000 chr1-0182 -0.086
183 chr1 15550001 15700000 chr1-0183 -0.125
184 chr1 15700001 15750000 chr1-0184 -0.213
185 chr1 15750001 15850000 chr1-0185 -0.149
186 chr1 15850001 15900000 chr1-0186 -0.216
187 chr1 15900001 15950000 chr1-0187 -0.123
188 chr1 15950001 16050000 chr1-0188 -0.195
189 chr1 16050001 16100000 chr1-0189 -0.073
190 chr1 16100001 16250000 chr1-0190 -0.141
191 chr1 16250001 16300000 chr1-0191 -0.082
192 chr1 16300001 16350000 chr1-0192 -0.134
193 chr1 16350001 16400000 chr1-0193 -0.072
194 chr1 16400001 16450000 chr1-0194 -0.106
195 chr1 16450001 16500000 chr1-0195 -0.133
196 chr1 16500001 16550000 chr1-0196 -0.060
197 chr1 16550001 16600000 chr1-0197 -0.116
198 chr1 16600001 16700000 chr1-0198 -0.150
199 chr1 16700001 16800000 chr1-0199 -0.145
200 chr1 16800001 16850000 chr1-0200 -0.095
201 chr1 16850001 16900000 chr1-0201 -0.107
202 chr1 16900001 16950000 chr1-0202 -0.131
203 chr1 16950001 17000000 chr1-0203 -0.139
204 chr1 17000001 17050000 chr1-0204 -0.023
205 chr1 17050001 17125658 chr1-0205 -0.066
206 chr1 17175659 17200000 chr1-0206 -0.030
207 chr1 17200001 17250000 chr1-0207 -0.048
208 chr1 17250001 17300000 chr1-0208 -0.078
209 chr1 17300001 17350000 chr1-0209 -0.081
210 chr1 17350001 17400000 chr1-0210 -0.059
211 chr1 17400001 17500000 chr1-0211 -0.144
212 chr1 17500001 17550000 chr1-0212 -0.142
213 chr1 17550001 17600000 chr1-0213 -0.201
214 chr1 17600001 17700000 chr1-0214 -0.159
215 chr1 17700001 17750000 chr1-0215 -0.146
216 chr1 17750001 17800000 chr1-0216 -0.105
217 chr1 17800001 17900000 chr1-0217 -0.115
218 chr1 17900001 17950000 chr1-0218 -0.071
219 chr1 17950001 18050000 chr1-0219 -0.101
220 chr1 18050001 18400000 chr1-0220 -0.175
221 chr1 18400001 18450000 chr1-0221 -0.123
222 chr1 18450001 18650000 chr1-0222 -0.131
223 chr1 18650001 18800000 chr1-0223 -0.182
224 chr1 18800001 18950000 chr1-0224 -0.148
225 chr1 18950001 19000000 chr1-0225 -0.064
226 chr1 19000001 19050000 chr1-0226 -0.206
227 chr1 19050001 19150000 chr1-0227 -0.116
228 chr1 19150001 19200000 chr1-0228 -0.151
229 chr1 19200001 19250000 chr1-0229 -0.079
230 chr1 19250001 19300000 chr1-0230 -0.099
231 chr1 19300001 19500000 chr1-0231 -0.117
232 chr1 19500001 19550000 chr1-0232 -0.192
233 chr1 19550001 19600000 chr1-0233 -0.147
234 chr1 19600001 19650000 chr1-0234 -0.168
235 chr1 19650001 19700000 chr1-0235 -0.102
236 chr1 19700001 19800000 chr1-0236 -0.224
237 chr1 19800001 19900000 chr1-0237 -0.118
238 chr1 19900001 19950000 chr1-0238 -0.142
239 chr1 19950001 20000000 chr1-0239 -0.085
240 chr1 20000001 20100000 chr1-0240 -0.148
241 chr1 20100001 20150000 chr1-0241 -0.156
242 chr1 20150001 20250000 chr1-0242 -0.119
243 chr1 20250001 20350000 chr1-0243 -0.168
244 chr1 20350001 20450000 chr1-0244 -0.185
245 chr1 20450001 20500000 chr1-0245 -0.113
246 chr1 20500001 20550000 chr1-0246 -0.174
247 chr1 20550001 20650000 chr1-0247 -0.155
248 chr1 20650001 20700000 chr1-0248 -0.112
249 chr1 20700001 20800000 chr1-0249 -0.133
250 chr1 20800001 20850000 chr1-0250 -0.121
251 chr1 20850001 20950000 chr1-0251 -0.030
252 chr1 20950001 21000000 chr1-0252 -0.135
253 chr1 21000001 21050000 chr1-0253 -0.075
254 chr1 21050001 21100000 chr1-0254 -0.106
255 chr1 21100001 21500000 chr1-0255 -0.151
256 chr1 21500001 21600000 chr1-0256 -0.157
257 chr1 21600001 21650000 chr1-0257 -0.116
258 chr1 21650001 21750000 chr1-0258 -0.129
259 chr1 21750001 21800000 chr1-0259 -0.040
260 chr1 21800001 21850000 chr1-0260 -0.065
261 chr1 21850001 21900000 chr1-0261 -0.228
262 chr1 21900001 21950000 chr1-0262 -0.103
263 chr1 21950001 22100000 chr1-0263 -0.164
264 chr1 22100001 22150000 chr1-0264 -0.068
265 chr1 22150001 22200000 chr1-0265 -0.091
266 chr1 22200001 22250000 chr1-0266 -0.095
267 chr1 22250001 22350000 chr1-0267 -0.167
268 chr1 22350001 22400000 chr1-0268 -0.178
269 chr1 22400001 22450000 chr1-0269 -0.139
270 chr1 22450001 22500000 chr1-0270 -0.120
271 chr1 22500001 22650000 chr1-0271 -0.110
272 chr1 22650001 22800000 chr1-0272 -0.138
273 chr1 22800001 22900000 chr1-0273 -0.057
274 chr1 22900001 22950000 chr1-0274 -0.097
275 chr1 22950001 23000000 chr1-0275 -0.074
276 chr1 23000001 23050000 chr1-0276 -0.145
277 chr1 23050001 23150000 chr1-0277 -0.106
278 chr1 23150001 23300000 chr1-0278 -0.087
279 chr1 23300001 23450000 chr1-0279 -0.124
280 chr1 23450001 23500000 chr1-0280 -0.268
281 chr1 23500001 23650000 chr1-0281 -0.138
282 chr1 23650001 23700000 chr1-0282 -0.156
283 chr1 23700001 23800000 chr1-0283 -0.139
284 chr1 23800001 23850000 chr1-0284 -0.116
285 chr1 23850001 23900000 chr1-0285 -0.134
286 chr1 23900001 24050000 chr1-0286 -0.254
287 chr1 24050001 24100000 chr1-0287 -0.034
288 chr1 24100001 24150000 chr1-0288 -0.147
289 chr1 24150001 24200000 chr1-0289 -0.152
290 chr1 24200001 24250000 chr1-0290 -0.170
291 chr1 24250001 24350000 chr1-0291 -0.148
292 chr1 24350001 24500000 chr1-0292 -0.137
293 chr1 24500001 24600000 chr1-0293 -0.131
294 chr1 24600001 24700000 chr1-0294 -0.180
295 chr1 24700001 24750000 chr1-0295 -0.144
296 chr1 24750001 24850000 chr1-0296 -0.086
297 chr1 24850001 24950000 chr1-0297 -0.266
298 chr1 24950001 25050000 chr1-0298 -0.122
299 chr1 25050001 25100000 chr1-0299 -0.106
300 chr1 25100001 25200000 chr1-0300 -0.147
301 chr1 25200001 25250000 chr1-0301 -0.158
302 chr1 25250001 25300000 chr1-0302 -0.111
303 chr1 25300001 25550000 chr1-0303 -0.123
304 chr1 25550001 25600000 chr1-0304 -0.125
305 chr1 25600001 25750000 chr1-0305 -0.171
306 chr1 25750001 25850000 chr1-0306 -0.234
307 chr1 25850001 25900000 chr1-0307 -0.080
308 chr1 25900001 25950000 chr1-0308 -0.097
309 chr1 25950001 26100000 chr1-0309 -0.186
310 chr1 26100001 26150000 chr1-0310 -0.156
311 chr1 26150001 26200000 chr1-0311 -0.124
312 chr1 26200001 26250000 chr1-0312 -0.118
313 chr1 26250001 26350000 chr1-0313 -0.151
314 chr1 26350001 26400000 chr1-0314 -0.114
315 chr1 26400001 26450000 chr1-0315 -0.093
316 chr1 26450001 26500000 chr1-0316 -0.063
317 chr1 26500001 26550000 chr1-0317 -0.128
318 chr1 26550001 26600000 chr1-0318 -0.143
319 chr1 26600001 26650000 chr1-0319 -0.099
320 chr1 26650001 26700000 chr1-0320 -0.161
321 chr1 26700001 26750000 chr1-0321 -0.102
322 chr1 26750001 26800000 chr1-0322 -0.105
323 chr1 26800001 26900000 chr1-0323 -0.169
324 chr1 26900001 27100000 chr1-0324 -0.142
325 chr1 27100001 27150000 chr1-0325 -0.188
326 chr1 27150001 27200000 chr1-0326 -0.092
327 chr1 27200001 27250000 chr1-0327 -0.168
328 chr1 27250001 27300000 chr1-0328 -0.095
329 chr1 27300001 27350000 chr1-0329 -0.136
330 chr1 27350001 27450000 chr1-0330 -0.077
331 chr1 27450001 27500000 chr1-0331 -0.149
332 chr1 27500001 27600000 chr1-0332 -0.131
333 chr1 27600001 27650000 chr1-0333 -0.106
334 chr1 27650001 27700000 chr1-0334 -0.090
335 chr1 27700001 27750000 chr1-0335 -0.119
336 chr1 27750001 27850000 chr1-0336 -0.139
337 chr1 27850001 27900000 chr1-0337 -0.093
338 chr1 27900001 27950000 chr1-0338 -0.133
339 chr1 27950001 28000000 chr1-0339 -0.165
340 chr1 28000001 28150000 chr1-0340 -0.128
341 chr1 28150001 28200000 chr1-0341 -0.119
342 chr1 28200001 28250000 chr1-0342 -0.121
343 chr1 28250001 28400000 chr1-0343 -0.110
344 chr1 28400001 28450000 chr1-0344 -0.182
345 chr1 28450001 28550000 chr1-0345 -0.207
346 chr1 28550001 28600000 chr1-0346 -0.183
347 chr1 28600001 28800000 chr1-0347 -0.133
348 chr1 28800001 28850000 chr1-0348 -0.201
349 chr1 28850001 28900000 chr1-0349 -0.126
350 chr1 28900001 28950000 chr1-0350 -0.104
351 chr1 28950001 29000000 chr1-0351
|