Supported by Dr. Osamu Ogasawara and  providing providing  . . |
|
Last data update: 2014.03.03 |
DecompositionDescriptionEven though the function Usage
proprius(y, X, type, offset = NULL, group = NULL,
mu = NULL, phi = NULL,
alpha = NULL, perm = 1000, plot = TRUE)
Arguments
DetailsThe user can provide a common The user can provide the confounding variable ValueIf ReferencesA Rauschenberger, MA Jonker, MA van de Wiel, and RX Menezes (2016). "Testing for association between RNA-Seq and high-dimensional data", BMC Bioinformatics. 17:118. html pdf (open access) JJ Goeman, SA van de Geer, F de Kort, and HC van Houwelingen (2004). "A global test for groups of genes: testing association with a clinical outcome", Bioinformatics. 20:93-99. html pdf (open access) See AlsoThe function Examples# simulate high-dimensional data n <- 30; p <- 100 y <- rnbinom(n,mu=10,size=1/0.25) X <- matrix(rnorm(n*p),nrow=n,ncol=p) # decomposition proprius(y,X,type="samples") proprius(y,X,type="covariates") Results
R version 3.3.1 (2016-06-21) -- "Bug in Your Hair"
Copyright (C) 2016 The R Foundation for Statistical Computing
Platform: x86_64-pc-linux-gnu (64-bit)
R is free software and comes with ABSOLUTELY NO WARRANTY.
You are welcome to redistribute it under certain conditions.
Type 'license()' or 'licence()' for distribution details.
R is a collaborative project with many contributors.
Type 'contributors()' for more information and
'citation()' on how to cite R or R packages in publications.
Type 'demo()' for some demos, 'help()' for on-line help, or
'help.start()' for an HTML browser interface to help.
Type 'q()' to quit R.
> library(globalSeq)
> png(filename="/home/ddbj/snapshot/RGM3/R_BC/result/globalSeq/proprius.Rd_%03d_medium.png", width=480, height=480)
> ### Name: proprius
> ### Title: Decomposition
> ### Aliases: proprius
> ### Keywords: methods
>
> ### ** Examples
>
>
> # simulate high-dimensional data
> n <- 30; p <- 100
> y <- rnbinom(n,mu=10,size=1/0.25)
> X <- matrix(rnorm(n*p),nrow=n,ncol=p)
>
> # decomposition
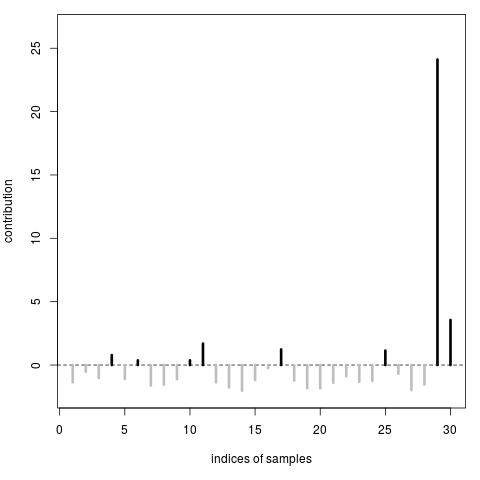
> proprius(y,X,type="samples")
[1] -1.3567212 -0.5201458 -1.0126671 0.7870157 -1.1039208 0.3675273
[7] -1.6114741 -1.5498190 -1.1134177 0.3722417 1.6947976 -1.3683284
[13] -1.7695083 -2.0365125 -1.1901664 -0.2313564 1.2302413 -1.2285466
[19] -1.8233707 -1.8346955 -1.3945066 -0.8868730 -1.3096920 -1.2489024
[25] 1.1394360 -0.6869264 -1.9868579 -1.5406372 24.1076220 3.5542340
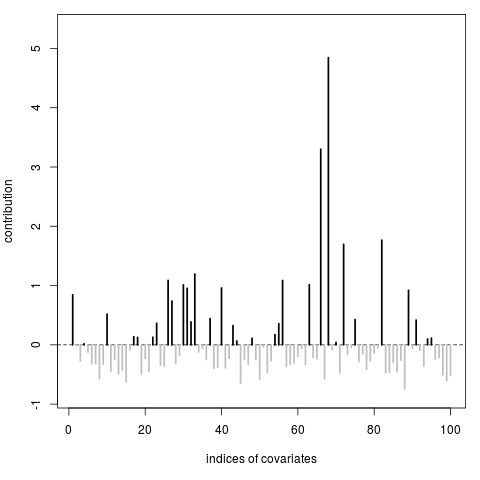
> proprius(y,X,type="covariates")
[1] 0.84550018 -0.01513445 -0.27117459 0.01821597 -0.12300187 -0.31693631
[7] -0.32056413 -0.57318360 -0.32912088 0.51803521 -0.43921767 -0.24243664
[13] -0.48524058 -0.42533687 -0.62583602 -0.08437057 0.13610674 0.12563903
[19] -0.49231860 -0.23040041 -0.45424874 0.12828120 0.36343064 -0.34336870
[25] -0.35957062 1.08730627 0.73845091 -0.31464951 -0.17449195 1.01096262
[31] 0.95386152 0.38579172 1.19353649 -0.12029903 -0.06347265 -0.24744725
[37] 0.44277813 -0.39403825 -0.37817400 0.95925079 -0.38768200 -0.22611421
[43] 0.32439396 0.06564774 -0.64392345 -0.24761051 -0.32846393 0.11069493
[49] -0.24770544 -0.57788636 -0.03804907 -0.47142164 -0.26815831 0.17231223
[55] 0.35719293 1.08604523 -0.36095299 -0.32494913 -0.30752131 -0.19480072
[61] -0.06466868 -0.33219776 1.01368264 -0.21156284 -0.23820781 3.30012922
[67] -0.57396646 4.84411670 -0.07969188 0.04110268 -0.47712917 1.69586457
[73] -0.15856122 -0.04271274 0.42774209 -0.27587226 -0.14833885 -0.41330645
[79] -0.27033254 -0.13942205 -0.04957926 1.76549609 -0.46838201 -0.46512151
[85] -0.29554955 -0.45505308 -0.25772613 -0.74533811 0.91903469 -0.05575209
[91] 0.41640106 -0.09516932 -0.35939054 0.09766040 0.11255990 -0.24257242
[97] -0.21531603 -0.50589800 -0.60256936 -0.52049407
>
>
>
>
>
>
> dev.off()
null device
1
>
|