Supported by Dr. Osamu Ogasawara and  providing providing  . . |
|
Last data update: 2014.03.03 |
Class "scan"DescriptionResults from scanning the genome with the inversion model for trial segments of fixed window size. Objects from the Class
Slots
Methods
Author(s)Alejandro Caceres acaceres@creal.cat See Also
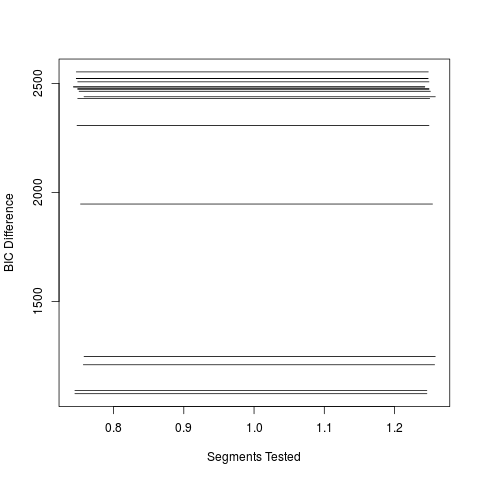
Examplesdata(scanRes) scanRes plot(scanRes,which="bic",thBic=0) Results
R version 3.3.1 (2016-06-21) -- "Bug in Your Hair"
Copyright (C) 2016 The R Foundation for Statistical Computing
Platform: x86_64-pc-linux-gnu (64-bit)
R is free software and comes with ABSOLUTELY NO WARRANTY.
You are welcome to redistribute it under certain conditions.
Type 'license()' or 'licence()' for distribution details.
R is a collaborative project with many contributors.
Type 'contributors()' for more information and
'citation()' on how to cite R or R packages in publications.
Type 'demo()' for some demos, 'help()' for on-line help, or
'help.start()' for an HTML browser interface to help.
Type 'q()' to quit R.
> library(inveRsion)
Loading required package: haplo.stats
Hola!
welcome to inevRsion package.
type: manual() for full manual
vignette("inveRsion") for a quick start
> png(filename="/home/ddbj/snapshot/RGM3/R_BC/result/inveRsion/scan-class.Rd_%03d_medium.png", width=480, height=480)
> ### Name: scan-class
> ### Title: Class "scan"
> ### Aliases: scan-class plot,scan-method show,scan-method
> ### Keywords: classes
>
> ### ** Examples
>
> data(scanRes)
> scanRes
-Showing object of class: scan-
Top 10 brake-points with highest Likelihood ratio:
LeftBP RightBP LogLike Prob BicDiff
1 0.75075-0.75124 1.25132-1.25160 2714.85 0.577 2464.021
2 0.74661-0.74661 1.24666-1.24687 2712.43 0.586 2522.406
3 0.74661-0.74690 1.24666-1.24687 2712.43 0.586 2522.406
4 0.74690-0.74721 1.24760-1.24881 2712.43 0.586 2522.406
5 0.74721-0.74776 1.24760-1.24881 2712.43 0.586 2552.810
6 0.74852-0.74890 1.24886-1.24911 2712.43 0.586 2507.204
7 0.74890-0.74891 1.24911-1.24930 2712.43 0.586 2476.800
8 0.74944-0.75066 1.24951-1.24951 2712.43 0.586 2431.195
9 0.75772-0.75789 1.25803-1.25821 2712.43 0.586 2438.796
10 0.74891-0.74944 1.24911-1.24930 2708.76 0.581 2473.131
others:
window: length window ( 0.5 ) for searching inversion segments
> plot(scanRes,which="bic",thBic=0)
>
>
>
>
>
> dev.off()
null device
1
>
|