Supported by Dr. Osamu Ogasawara and  providing providing  . . |
|
Last data update: 2014.03.03 |
Estimate the copy number profileDescriptionFunction to estimate the copy number profile with a piecewise constant function using mBPCR. Eventually, it is possible to estimate the profile with a
smoothing curve using either the Bayesian Regression Curve with K_2 (BRC with K_2) or the Bayesian Regression Curve Averaging over k (BRCAk). It is also possible
to choose the estimator of the variance of the levels UsagecomputeMBPCR(y, kMax=50, nu=NULL, rhoSquare=NULL, sigmaSquare=NULL, typeEstRho=1, regr=NULL) Arguments
DetailsBy default, the function estimates the copy number profile with mBPCR and estimating rhoSquare on the sample, using hat{ρ}_1^2. It is
also possible to use hat{ρ}^2 as estimator of The function gives also the possibility to estimate the profile with a Bayesian regression curve: if ValueA list containing:
ReferencesRancoita, P. M. V., Hutter, M., Bertoni, F., Kwee, I. (2009). Bayesian DNA copy number analysis. BMC Bioinformatics 10: 10. http://www.idsia.ch/~paola/mBPCR See Also
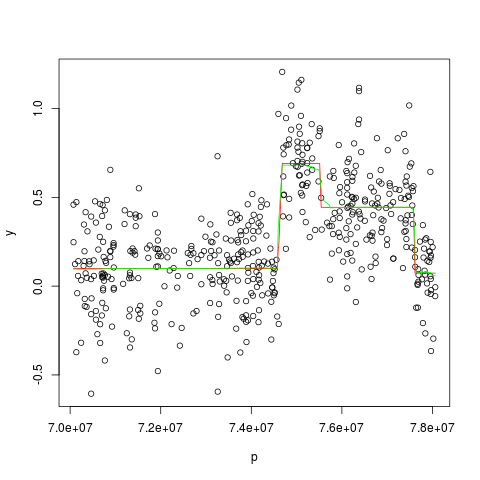
Examples##import the 250K NSP data of chromosome 11 of cell line JEKO-1 data(jekoChr11Array250Knsp) ##first example ## we select a part of chromosome 11 y <- jekoChr11Array250Knsp$log2ratio[6400:6900] p <- jekoChr11Array250Knsp$PhysicalPosition[6400:6900] ##we estimate the profile using the global parameters estimated on the whole genome ##the profile is estimated with mBPCR and with the Bayesian Regression Curve results <- computeMBPCR(y, nu=-3.012772e-10, rhoSquare=0.0479, sigmaSquare=0.0699, regr="BRC") plot(p, y) points(p, results$estPC, type='l', col='red') points(p, results$regrCurve,type='l', col='green') ###second example ### we select a part of chromosome 11 #y <- jekoChr11Array250Knsp$log2ratio[10600:11600] #p <- jekoChr11Array250Knsp$PhysicalPosition[10600:11600] ###we estimate the profile using the global parameters estimated on the whole genome ###the profile is estimated with mBPCR and with the Bayesian Regression Curve Ak #results <- computeMBPCR(y, nu=-3.012772e-10, rhoSquare=0.0479, sigmaSquare=0.0699, regr="BRCAk") #plot(p,y) #points(p, results$estPC, type='l', col='red') #points(p, results$regrCurve, type='l', col='green') Results
R version 3.3.1 (2016-06-21) -- "Bug in Your Hair"
Copyright (C) 2016 The R Foundation for Statistical Computing
Platform: x86_64-pc-linux-gnu (64-bit)
R is free software and comes with ABSOLUTELY NO WARRANTY.
You are welcome to redistribute it under certain conditions.
Type 'license()' or 'licence()' for distribution details.
R is a collaborative project with many contributors.
Type 'contributors()' for more information and
'citation()' on how to cite R or R packages in publications.
Type 'demo()' for some demos, 'help()' for on-line help, or
'help.start()' for an HTML browser interface to help.
Type 'q()' to quit R.
> library(mBPCR)
Loading required package: oligoClasses
Welcome to oligoClasses version 1.34.0
Loading required package: SNPchip
Welcome to SNPchip version 2.18.0
> png(filename="/home/ddbj/snapshot/RGM3/R_BC/result/mBPCR/computeMBPCR.Rd_%03d_medium.png", width=480, height=480)
> ### Name: computeMBPCR
> ### Title: Estimate the copy number profile
> ### Aliases: computeMBPCR
> ### Keywords: regression smooth
>
> ### ** Examples
>
> ##import the 250K NSP data of chromosome 11 of cell line JEKO-1
> data(jekoChr11Array250Knsp)
>
>
> ##first example
> ## we select a part of chromosome 11
> y <- jekoChr11Array250Knsp$log2ratio[6400:6900]
> p <- jekoChr11Array250Knsp$PhysicalPosition[6400:6900]
> ##we estimate the profile using the global parameters estimated on the whole genome
> ##the profile is estimated with mBPCR and with the Bayesian Regression Curve
> results <- computeMBPCR(y, nu=-3.012772e-10, rhoSquare=0.0479, sigmaSquare=0.0699, regr="BRC")
Computation of log(A^0)
Computation of left and right recursions
Determination of PC Regression
Computation of Bayesian regression Curve
> plot(p, y)
> points(p, results$estPC, type='l', col='red')
> points(p, results$regrCurve,type='l', col='green')
>
> ###second example
> ### we select a part of chromosome 11
> #y <- jekoChr11Array250Knsp$log2ratio[10600:11600]
> #p <- jekoChr11Array250Knsp$PhysicalPosition[10600:11600]
> ###we estimate the profile using the global parameters estimated on the whole genome
> ###the profile is estimated with mBPCR and with the Bayesian Regression Curve Ak
> #results <- computeMBPCR(y, nu=-3.012772e-10, rhoSquare=0.0479, sigmaSquare=0.0699, regr="BRCAk")
> #plot(p,y)
> #points(p, results$estPC, type='l', col='red')
> #points(p, results$regrCurve, type='l', col='green')
>
>
>
>
>
>
> dev.off()
null device
1
>
|