Supported by Dr. Osamu Ogasawara and  providing providing  . . |
|
Last data update: 2014.03.03 |
Class "marrayNorm", classes and methods for post-normalization cDNA microarray intensity dataDescriptionThis class represents post-normalization intensity data for a batch of cDNA microarrays. A batch of arrays consists of a collection of arrays with the same layout ( Objects from the ClassObjects can be created by calls of the form
Slots
Methods
Author(s)Sandrine Dudoit, http://www.stat.berkeley.edu/~sandrine. ReferencesS. Dudoit and Y. H. Yang. (2002). Bioconductor R packages for exploratory analysis and normalization of cDNA microarray data. In G. Parmigiani, E. S. Garrett, R. A. Irizarry and S. L. Zeger, editors, The Analysis of Gene Expression Data: Methods and Software, Springer, New York. See Also
Examples
# Examples use swirl dataset, for description type ? swirl
data(swirl)
# Median normalization
mnorm<-maNorm(swirl[,2:3],norm="m")
# Object of class marrayNorm for the second and third swirl arrays
mnorm
# Function call
maNormCall(mnorm)
# Object of class marrayInfo -- Probe sequences
maGnames(mnorm)
# Object of class marrayInfo -- Target samples
maTargets(mnorm)
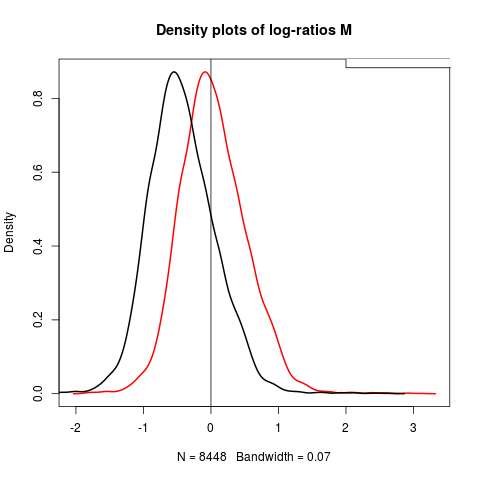
# Density plot of log-ratios M for third array
plot(density(maM(mnorm[,2])), lwd=2, col=2, main="Density plots of log-ratios M")
lines(density(maM(swirl[,3])), lwd=2)
abline(v=0)
legend(2,1,c("Pre-normalization","Post-normalization"))
Results
R version 3.3.1 (2016-06-21) -- "Bug in Your Hair"
Copyright (C) 2016 The R Foundation for Statistical Computing
Platform: x86_64-pc-linux-gnu (64-bit)
R is free software and comes with ABSOLUTELY NO WARRANTY.
You are welcome to redistribute it under certain conditions.
Type 'license()' or 'licence()' for distribution details.
R is a collaborative project with many contributors.
Type 'contributors()' for more information and
'citation()' on how to cite R or R packages in publications.
Type 'demo()' for some demos, 'help()' for on-line help, or
'help.start()' for an HTML browser interface to help.
Type 'q()' to quit R.
> library(marray)
Loading required package: limma
> png(filename="/home/ddbj/snapshot/RGM3/R_BC/result/marray/marrayNorm-class.Rd_%03d_medium.png", width=480, height=480)
> ### Name: marrayNorm-class
> ### Title: Class "marrayNorm", classes and methods for post-normalization
> ### cDNA microarray intensity data
> ### Aliases: marrayNorm-class marrayNorm maA maA<- maM maM<- maMloc
> ### maMloc<- maMscale maMscale<- [,marrayNorm-method
> ### cbind,marrayNorm-method coerce,marrayRaw,marrayNorm-method
> ### maA<-,marrayNorm,matrix-method maA,marrayNorm-method
> ### maControls<-,marrayNorm-method maControls,marrayNorm-method
> ### maGnames,marrayNorm-method maGridCol,marrayNorm-method
> ### maGridRow,marrayNorm-method maLayout,marrayNorm-method
> ### maMloc<-,marrayNorm,matrix-method maMloc,marrayNorm-method
> ### maM<-,marrayNorm,matrix-method maM,marrayNorm-method
> ### maMscale<-,marrayNorm,matrix-method maMscale,marrayNorm-method
> ### maNgc<-,marrayNorm,numeric-method maNgc,marrayNorm-method
> ### maNgr<-,marrayNorm,numeric-method maNgr,marrayNorm-method
> ### maNormCall,marrayNorm-method maNotes<-,marrayNorm,character-method
> ### maNotes,marrayNorm-method maNsamples,marrayNorm-method
> ### maNsc<-,marrayNorm,numeric-method maNsc,marrayNorm-method
> ### maNspots<-,marrayNorm,numeric-method maNspots,marrayNorm-method
> ### maNsr<-,marrayNorm,numeric-method maNsr,marrayNorm-method
> ### maPlate<-,marrayNorm-method maPlate,marrayNorm-method
> ### maPrintTip,marrayNorm-method maSpotCol,marrayNorm-method
> ### maSpotRow,marrayNorm-method maSub<-,marrayNorm-method
> ### maSub,marrayNorm-method maTargets,marrayNorm-method
> ### maW<-,marrayNorm,matrix-method maW,marrayNorm-method
> ### maLG,marrayNorm-method maLR,marrayNorm-method print,marrayNorm-method
> ### show,marrayNorm-method summary,marrayNorm-method maNormCall
> ### Keywords: classes
>
> ### ** Examples
>
> # Examples use swirl dataset, for description type ? swirl
>
> data(swirl)
>
> # Median normalization
> mnorm<-maNorm(swirl[,2:3],norm="m")
>
> # Object of class marrayNorm for the second and third swirl arrays
> mnorm
An object of class "marrayNorm"
@maA
C:/GNU/R/R-2.4.1/library/marray/swirldata/swirl.2.spot
[1,] 14.09380
[2,] 14.21926
[3,] 14.18555
[4,] 12.69088
[5,] 12.59373
C:/GNU/R/R-2.4.1/library/marray/swirldata/swirl.3.spot
[1,] 11.412575
[2,] 11.449228
[3,] 11.282760
[4,] 8.148732
[5,] 9.594076
8443 more rows ...
@maM
C:/GNU/R/R-2.4.1/library/marray/swirldata/swirl.2.spot
[1,] -0.285832418
[2,] -0.342926131
[3,] -0.263816797
[4,] 0.003256647
[5,] -0.145964826
C:/GNU/R/R-2.4.1/library/marray/swirldata/swirl.3.spot
[1,] 0.5510083
[2,] 0.5600710
[3,] 0.6486954
[4,] 0.1846651
[5,] 0.2544244
8443 more rows ...
@maMloc
[,1] [,2]
[1,] 0.03029219 -0.4602068
[2,] 0.03029219 -0.4602068
[3,] 0.03029219 -0.4602068
[4,] 0.03029219 -0.4602068
[5,] 0.03029219 -0.4602068
8443 more rows ...
@maMscale
<0 x 0 matrix>
@maW
<0 x 0 matrix>
@maLayout
An object of class "marrayLayout"
@maNgr
[1] 4
@maNgc
[1] 4
@maNsr
[1] 22
@maNsc
[1] 24
@maNspots
[1] 8448
@maSub
[1] TRUE TRUE TRUE TRUE TRUE
8443 more elements ...
@maPlate
[1] 1 1 1 1 1
Levels: 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22
8443 more elements ...
@maControls
[1] 1 1 1 1 1
Levels: 0 1
8443 more elements ...
@maNotes
[1] "No Input File"
@maGnames
An object of class "marrayInfo"
@maLabels
[1] "geno1" "geno2" "geno3" "3XSSC" "3XSSC"
8443 more elements ...
@maInfo
"ID" "Name"
1 control geno1
2 control geno2
3 control geno3
4 control 3XSSC
5 control 3XSSC
8443 more rows ...
@maNotes
[1] "C:/GNU/R/R-2.4.1/library/marray/swirldata/fish.gal"
@maTargets
An object of class "marrayInfo"
@maLabels
[1] "swirl.2.spot" "swirl.3.spot"
@maInfo
Names slide number experiment Cy3 experiment Cy5 date comments
2 swirl.2.spot 82 wild type swirl 2001/9/20 NA
3 swirl.3.spot 93 swirl wild type 2001/11/8 NA
@maNotes
[1] "C:/GNU/R/R-2.4.1/library/marray/swirldata/SwirlSample.txt"
@maNotes
[1] "Spot Data"
@maNormCall
maNormMain(mbatch = mbatch, f.loc = list(maNormMed(x = NULL,
y = "maM", subset = subset)), Mloc = Mloc, Mscale = Mscale,
echo = echo)
>
> # Function call
> maNormCall(mnorm)
maNormMain(mbatch = mbatch, f.loc = list(maNormMed(x = NULL,
y = "maM", subset = subset)), Mloc = Mloc, Mscale = Mscale,
echo = echo)
>
> # Object of class marrayInfo -- Probe sequences
> maGnames(mnorm)
An object of class "marrayInfo"
@maLabels
[1] "geno1" "geno2" "geno3" "3XSSC" "3XSSC"
8443 more elements ...
@maInfo
"ID" "Name"
1 control geno1
2 control geno2
3 control geno3
4 control 3XSSC
5 control 3XSSC
8443 more rows ...
@maNotes
[1] "C:/GNU/R/R-2.4.1/library/marray/swirldata/fish.gal"
>
> # Object of class marrayInfo -- Target samples
> maTargets(mnorm)
An object of class "marrayInfo"
@maLabels
[1] "swirl.2.spot" "swirl.3.spot"
@maInfo
Names slide number experiment Cy3 experiment Cy5 date comments
2 swirl.2.spot 82 wild type swirl 2001/9/20 NA
3 swirl.3.spot 93 swirl wild type 2001/11/8 NA
@maNotes
[1] "C:/GNU/R/R-2.4.1/library/marray/swirldata/SwirlSample.txt"
>
> # Density plot of log-ratios M for third array
> plot(density(maM(mnorm[,2])), lwd=2, col=2, main="Density plots of log-ratios M")
> lines(density(maM(swirl[,3])), lwd=2)
> abline(v=0)
> legend(2,1,c("Pre-normalization","Post-normalization"))
>
>
>
>
>
> dev.off()
null device
1
>
|