Supported by Dr. Osamu Ogasawara and  providing providing  . . |
|
Last data update: 2014.03.03 |
Class "marrayRaw", classes and methods for pre-normalization cDNA microarray intensity dataDescriptionThis class represents pre-normalization intensity data for
a batch of cDNA microarrays. A batch of arrays consists of a
collection of arrays with the same layout
( Objects from the ClassObjects can be created by calls of the form
Slots
Methods
Author(s)Sandrine Dudoit, http://www.stat.berkeley.edu/~sandrine. ReferencesS. Dudoit and Y. H. Yang. (2002). Bioconductor R packages for exploratory analysis and normalization of cDNA microarray data. In G. Parmigiani, E. S. Garrett, R. A. Irizarry and S. L. Zeger, editors, The Analysis of Gene Expression Data: Methods and Software, Springer, New York. See Also
Examples# Examples use swirl dataset, for description type ? swirl require(limma) data(swirl) # Object of class marrayRaw for the 4 swirl arrays swirl # Object of class marrayLayout maLayout(swirl) # Access only the first 100 spots of the third array swirl[1:100,3] # Accessor methods -- How many spots on the array maNspots(swirl) # Density plot of log-ratios M for third array plot(density(maM(swirl[,3]))) # Assignment methods -- Replace maNotes slot maNotes(swirl) maNotes(swirl)<-"This is a zebrafish microarray" maNotes(swirl) Results
R version 3.3.1 (2016-06-21) -- "Bug in Your Hair"
Copyright (C) 2016 The R Foundation for Statistical Computing
Platform: x86_64-pc-linux-gnu (64-bit)
R is free software and comes with ABSOLUTELY NO WARRANTY.
You are welcome to redistribute it under certain conditions.
Type 'license()' or 'licence()' for distribution details.
R is a collaborative project with many contributors.
Type 'contributors()' for more information and
'citation()' on how to cite R or R packages in publications.
Type 'demo()' for some demos, 'help()' for on-line help, or
'help.start()' for an HTML browser interface to help.
Type 'q()' to quit R.
> library(marray)
Loading required package: limma
> png(filename="/home/ddbj/snapshot/RGM3/R_BC/result/marray/marrayRaw-class.Rd_%03d_medium.png", width=480, height=480)
> ### Name: marrayRaw-class
> ### Title: Class "marrayRaw", classes and methods for pre-normalization
> ### cDNA microarray intensity data
> ### Aliases: marrayRaw-class marrayRaw maRf maRf<- maGf maGf<- maRb maRb<-
> ### maGb maGb<- maW maW<- maLayout maLayout<- [,marrayRaw-method
> ### cbind,marrayRaw-method maA,marrayRaw-method
> ### maControls<-,marrayRaw-method maControls,marrayRaw-method
> ### maGb<-,marrayRaw,matrix-method maGb<-,marrayRaw,NULL-method
> ### maGb,marrayRaw-method maGf<-,marrayRaw,matrix-method
> ### maGf,marrayRaw-method maGnames,marrayRaw-method
> ### maGridCol,marrayRaw-method maGridRow,marrayRaw-method
> ### maLayout,marrayRaw-method maLG,marrayRaw-method maLR,marrayRaw-method
> ### maM,marrayRaw-method maNgc<-,marrayRaw,numeric-method
> ### maNgc,marrayRaw-method maNgr<-,marrayRaw,numeric-method
> ### maNgr,marrayRaw-method maNotes<-,marrayRaw,character-method
> ### maNotes,marrayRaw-method maNsamples,marrayRaw-method
> ### maNsc<-,marrayRaw,numeric-method maNsc,marrayRaw-method
> ### maNspots<-,marrayRaw,numeric-method maNspots,marrayRaw-method
> ### maNsr<-,marrayRaw,numeric-method maNsr,marrayRaw-method
> ### maPlate<-,marrayRaw-method maPlate,marrayRaw-method
> ### maPrintTip,marrayRaw-method maRb<-,marrayRaw,matrix-method
> ### maRb<-,marrayRaw,NULL-method maRb,marrayRaw-method
> ### maRf<-,marrayRaw,matrix-method maRf,marrayRaw-method
> ### maSpotCol,marrayRaw-method maSpotRow,marrayRaw-method
> ### maSub<-,marrayRaw-method maSub,marrayRaw-method
> ### maTargets,marrayRaw-method maW<-,marrayRaw,matrix-method
> ### maW,marrayRaw-method print,marrayRaw-method show,marrayRaw-method
> ### summary,marrayRaw-method maGnames maGnames<- maTargets maTargets<-
> ### maLR maLG maNsamples
> ### Keywords: classes
>
> ### ** Examples
>
> # Examples use swirl dataset, for description type ? swirl
> require(limma)
> data(swirl)
>
> # Object of class marrayRaw for the 4 swirl arrays
> swirl
An object of class "marrayRaw"
@maRf
C:/GNU/R/R-2.4.1/library/marray/swirldata/swirl.1.spot
[1,] 19538.470
[2,] 23619.820
[3,] 21579.950
[4,] 8905.143
[5,] 8676.095
C:/GNU/R/R-2.4.1/library/marray/swirldata/swirl.2.spot
[1,] 16138.720
[2,] 17247.670
[3,] 17317.150
[4,] 6794.381
[5,] 6043.542
C:/GNU/R/R-2.4.1/library/marray/swirldata/swirl.3.spot
[1,] 2895.1600
[2,] 2976.6230
[3,] 2735.6190
[4,] 318.9524
[5,] 780.6667
C:/GNU/R/R-2.4.1/library/marray/swirldata/swirl.4.spot
[1,] 14054.5400
[2,] 20112.2600
[3,] 12945.8500
[4,] 524.0476
[5,] 304.6190
8443 more rows ...
@maGf
C:/GNU/R/R-2.4.1/library/marray/swirldata/swirl.1.spot
[1,] 22028.260
[2,] 25613.200
[3,] 22652.390
[4,] 8929.286
[5,] 8746.476
C:/GNU/R/R-2.4.1/library/marray/swirldata/swirl.2.spot
[1,] 19278.770
[2,] 21438.960
[3,] 20386.470
[4,] 6677.619
[5,] 6576.292
C:/GNU/R/R-2.4.1/library/marray/swirldata/swirl.3.spot
[1,] 2727.5600
[2,] 2787.0330
[3,] 2419.8810
[4,] 383.2381
[5,] 901.0000
C:/GNU/R/R-2.4.1/library/marray/swirldata/swirl.4.spot
[1,] 19930.6500
[2,] 25426.5800
[3,] 16225.9500
[4,] 786.9048
[5,] 468.0476
8443 more rows ...
@maRb
C:/GNU/R/R-2.4.1/library/marray/swirldata/swirl.1.spot
[1,] 174
[2,] 174
[3,] 174
[4,] 163
[5,] 140
C:/GNU/R/R-2.4.1/library/marray/swirldata/swirl.2.spot
[1,] 136
[2,] 133
[3,] 133
[4,] 105
[5,] 105
C:/GNU/R/R-2.4.1/library/marray/swirldata/swirl.3.spot
[1,] 82
[2,] 82
[3,] 76
[4,] 61
[5,] 61
C:/GNU/R/R-2.4.1/library/marray/swirldata/swirl.4.spot
[1,] 48
[2,] 48
[3,] 48
[4,] 48
[5,] 49
8443 more rows ...
@maGb
C:/GNU/R/R-2.4.1/library/marray/swirldata/swirl.1.spot
[1,] 182
[2,] 171
[3,] 153
[4,] 153
[5,] 153
C:/GNU/R/R-2.4.1/library/marray/swirldata/swirl.2.spot
[1,] 175
[2,] 183
[3,] 183
[4,] 142
[5,] 142
C:/GNU/R/R-2.4.1/library/marray/swirldata/swirl.3.spot
[1,] 86
[2,] 86
[3,] 86
[4,] 71
[5,] 71
C:/GNU/R/R-2.4.1/library/marray/swirldata/swirl.4.spot
[1,] 97
[2,] 85
[3,] 85
[4,] 87
[5,] 87
8443 more rows ...
@maW
<0 x 0 matrix>
@maLayout
An object of class "marrayLayout"
@maNgr
[1] 4
@maNgc
[1] 4
@maNsr
[1] 22
@maNsc
[1] 24
@maNspots
[1] 8448
@maSub
[1] TRUE
@maPlate
factor(0)
Levels:
@maControls
[1] 1 1 1 1 1
Levels: 0 1
8443 more elements ...
@maNotes
[1] "No Input File"
@maGnames
An object of class "marrayInfo"
@maLabels
[1] "geno1" "geno2" "geno3" "3XSSC" "3XSSC"
8443 more elements ...
@maInfo
"ID" "Name"
1 control geno1
2 control geno2
3 control geno3
4 control 3XSSC
5 control 3XSSC
8443 more rows ...
@maNotes
[1] "C:/GNU/R/R-2.4.1/library/marray/swirldata/fish.gal"
@maTargets
An object of class "marrayInfo"
@maLabels
[1] "swirl.1.spot" "swirl.2.spot" "swirl.3.spot" "swirl.4.spot"
@maInfo
Names slide number experiment Cy3 experiment Cy5 date comments
1 swirl.1.spot 81 swirl wild type 2001/9/20 NA
2 swirl.2.spot 82 wild type swirl 2001/9/20 NA
3 swirl.3.spot 93 swirl wild type 2001/11/8 NA
4 swirl.4.spot 94 wild type swirl 2001/11/8 NA
@maNotes
[1] "C:/GNU/R/R-2.4.1/library/marray/swirldata/SwirlSample.txt"
@maNotes
[1] "Spot Data"
>
> # Object of class marrayLayout
> maLayout(swirl)
An object of class "marrayLayout"
@maNgr
[1] 4
@maNgc
[1] 4
@maNsr
[1] 22
@maNsc
[1] 24
@maNspots
[1] 8448
@maSub
[1] TRUE
@maPlate
factor(0)
Levels:
@maControls
[1] 1 1 1 1 1
Levels: 0 1
8443 more elements ...
@maNotes
[1] "No Input File"
>
> # Access only the first 100 spots of the third array
> swirl[1:100,3]
An object of class "marrayRaw"
@maRf
[1] 2895.1600 2976.6230 2735.6190 318.9524 780.6667
95 more rows ...
@maGf
[1] 2727.5600 2787.0330 2419.8810 383.2381 901.0000
95 more rows ...
@maRb
[1] 82 82 76 61 61
95 more rows ...
@maGb
[1] 86 86 86 71 71
95 more rows ...
@maW
<0 x 0 matrix>
@maLayout
An object of class "marrayLayout"
@maNgr
[1] 4
@maNgc
[1] 4
@maNsr
[1] 22
@maNsc
[1] 24
@maNspots
[1] 8448
@maSub
[1] TRUE TRUE TRUE TRUE TRUE
8443 more elements ...
@maPlate
[1] 1 1 1 1 1
Levels: 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22
95 more elements ...
@maControls
[1] 1 1 1 1 1
Levels: 0 1
95 more elements ...
@maNotes
[1] "No Input File"
@maGnames
An object of class "marrayInfo"
@maLabels
[1] "geno1" "geno2" "geno3" "3XSSC" "3XSSC"
95 more elements ...
@maInfo
"ID" "Name"
1 control geno1
2 control geno2
3 control geno3
4 control 3XSSC
5 control 3XSSC
95 more rows ...
@maNotes
[1] "C:/GNU/R/R-2.4.1/library/marray/swirldata/fish.gal"
@maTargets
An object of class "marrayInfo"
@maLabels
[1] "swirl.3.spot"
@maInfo
Names slide number experiment Cy3 experiment Cy5 date comments
3 swirl.3.spot 93 swirl wild type 2001/11/8 NA
@maNotes
[1] "C:/GNU/R/R-2.4.1/library/marray/swirldata/SwirlSample.txt"
@maNotes
[1] "Spot Data"
>
> # Accessor methods -- How many spots on the array
> maNspots(swirl)
[1] 8448
>
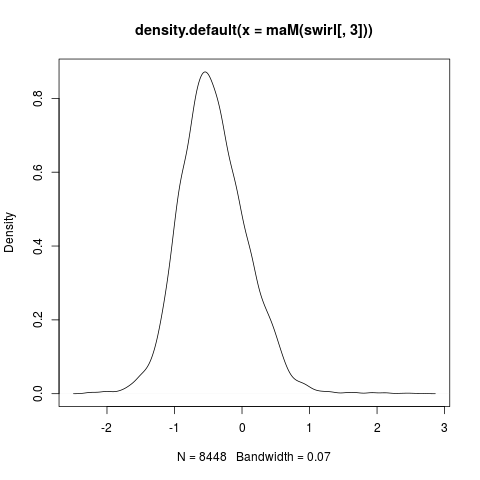
> # Density plot of log-ratios M for third array
> plot(density(maM(swirl[,3])))
>
> # Assignment methods -- Replace maNotes slot
> maNotes(swirl)
[1] "Spot Data"
> maNotes(swirl)<-"This is a zebrafish microarray"
> maNotes(swirl)
[1] "This is a zebrafish microarray"
>
>
>
>
>
> dev.off()
null device
1
>
|