Supported by Dr. Osamu Ogasawara and  providing providing  . . |
|
Last data update: 2014.03.03 |
The plot method for a MDQC objectDescriptionThe plot method for a MDQC object, which plots ... Usage## S3 method for class 'mdqc' plot(x, levels = c(0.9, 0.95, 0.99), xlab="", ylab="", mfrow=NULL, mfcol=NULL, ...) Arguments
DetailsThis plot method is for the output from the function
For further details, see Cohen Freue et al. (2007) Author(s)Justin Harrington harringt@stat.ubc.ca and Gabriela V. Cohen Freue gcohen@stat.ubc.ca. ReferencesCohen Freue, G. V. and Hollander, Z. and Shen, E. and Zamar, R. H. and Balshaw, R. and Scherer, A. and McManus, B. and Keown, P. and McMaster, W. R. and Ng, R. T. (2007) ‘MDQC: A New Quality Assessment Method for Microarrays Based on Quality Control Reports’. Bioinformatics 23, 3162 – 3169. See Also
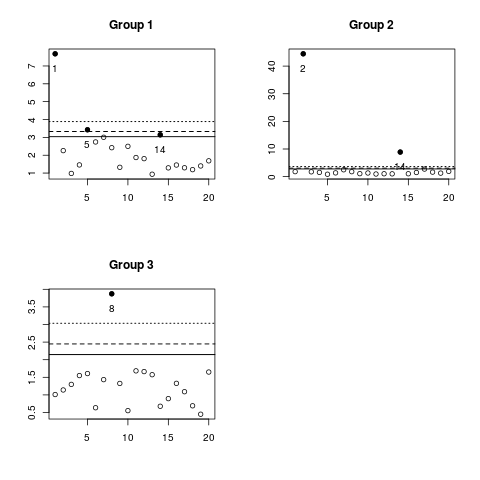
Examplesdata(allQC) mdout <- mdqc(allQC, method="cluster", k=3) plot(mdout) ## Just one critical value plot(mdout, levels=0.9) Results
R version 3.3.1 (2016-06-21) -- "Bug in Your Hair"
Copyright (C) 2016 The R Foundation for Statistical Computing
Platform: x86_64-pc-linux-gnu (64-bit)
R is free software and comes with ABSOLUTELY NO WARRANTY.
You are welcome to redistribute it under certain conditions.
Type 'license()' or 'licence()' for distribution details.
R is a collaborative project with many contributors.
Type 'contributors()' for more information and
'citation()' on how to cite R or R packages in publications.
Type 'demo()' for some demos, 'help()' for on-line help, or
'help.start()' for an HTML browser interface to help.
Type 'q()' to quit R.
> library(mdqc)
Loading required package: cluster
Loading required package: MASS
> png(filename="/home/ddbj/snapshot/RGM3/R_BC/result/mdqc/plot.mdqc.Rd_%03d_medium.png", width=480, height=480)
> ### Name: plot.mdqc
> ### Title: The plot method for a MDQC object
> ### Aliases: plot.mdqc
> ### Keywords: multivariate robust
>
> ### ** Examples
>
> data(allQC)
> mdout <- mdqc(allQC, method="cluster", k=3)
> plot(mdout)
>
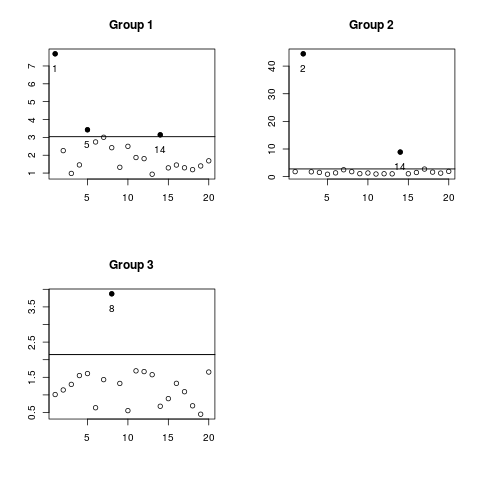
> ## Just one critical value
> plot(mdout, levels=0.9)
>
>
>
>
>
> dev.off()
null device
1
>
|