Supported by Dr. Osamu Ogasawara and  providing providing  . . |
|
Last data update: 2014.03.03 |
Find optimal prognostic features using the Messina algorithmDescriptionRun the MessinaSurv algorithm to find features (eg. genes) that can define groups of patients with very different survival times. UsagemessinaSurv(x, y, obj_min, obj_func, min_group_frac = 0.1, f_train = 0.8, n_boot = 50, seed = NULL, parallel = NULL, silent = FALSE) Arguments
DetailsThe MessinaSurv algorithm aims to identify features for which patients with high signal and patients with low signal have very different survival outcomes. This is achieved by definining an objective function which assigns a numerical value to how strongly the survival in two groups of patients differs, then assessing the value of this objective at different signal levels of each feature. Those features for which, at a given signal level, the objective function is consistently above a user-supplied minimum level, are selected by MessinaSurv as being single-feature survival predictors. MessinaSurv has applications as an algorithm to identify features that are survival-related, as well as a principled method to identify threshold signal values to separate a cohort into poor- and good-prognosis subgroups. It can also be used as a feature filter, selecting and discretising survival-related features before they are input into a multivariate predictor. Valuean object of class "MessinaSurvResult" containing the results of the analysis. Objective functionsMessinaSurv uses the value of its objective function as a measure of the strength of the difference in survival of the two patient groups defined by the threshold. Three objective functions are currently defined:
Methods "coxcoef" and "reltau" show instability for very high and low threshold values, and so should be used with an appropriate value of min_group_frac for stable fits. Method "tau" is stable to extreme threshold values, and therefore will tolerate min_group_frac = 0, however note that "tau" naturally penalizes small subgroups, and is therefore a poor choice unless you wish to find approximately equal-sized subgroups. Minimum group fractionThe parameter min_group_frac limits the size of the smallest subgroups that messinaSurv can select. As the groups become smaller, the "reltau" and "coxcoef" objective functions become unstable, and can generate spurious results. These are seen on the diagnostics produced by the messina plot functions as very high objective values at very low and high threshold values. To control these results, set min_group_frac to a high enough value that the objective functions reliably fit. Generally, max(0.1, 10/N), where N is the total number of patients, is sufficient. Keep in mind that setting this parameter too high will limit messinaSurv's ability to identify small subsets of patients with dramatically different survival from the rest: the smallest subset that will be reliably identified is min_group_frac of patients. Author(s)Mark Pinese m.pinese@garvan.org.au See Also
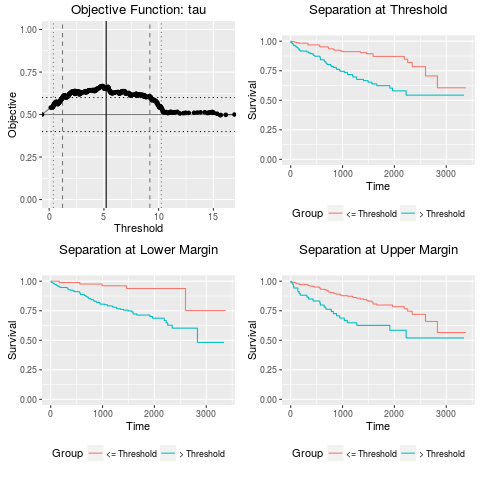
Examples## Load a subset of the TCGA renal clear cell carcinoma data ## as an example. data(tcga_kirc_example) ## Run the messinaSurv analysis on these data. Use a tau ## objective, with a minimum performance of 0.6. Note that ## messinaSurv analyses are very computationally-intensive, ## so in actual use multicore use with doMC and parallel = TRUE ## is strongly recommended. fit = messinaSurv(kirc.exprs, kirc.surv, obj_func = "tau", obj_min = 0.6) fit plot(fit) Results
R version 3.3.1 (2016-06-21) -- "Bug in Your Hair"
Copyright (C) 2016 The R Foundation for Statistical Computing
Platform: x86_64-pc-linux-gnu (64-bit)
R is free software and comes with ABSOLUTELY NO WARRANTY.
You are welcome to redistribute it under certain conditions.
Type 'license()' or 'licence()' for distribution details.
R is a collaborative project with many contributors.
Type 'contributors()' for more information and
'citation()' on how to cite R or R packages in publications.
Type 'demo()' for some demos, 'help()' for on-line help, or
'help.start()' for an HTML browser interface to help.
Type 'q()' to quit R.
> library(messina)
Loading required package: survival
> png(filename="/home/ddbj/snapshot/RGM3/R_BC/result/messina/messinaSurv.Rd_%03d_medium.png", width=480, height=480)
> ### Name: messinaSurv
> ### Title: Find optimal prognostic features using the Messina algorithm
> ### Aliases: messinaSurv
>
> ### ** Examples
>
> ## Load a subset of the TCGA renal clear cell carcinoma data
> ## as an example.
> data(tcga_kirc_example)
>
> ## Run the messinaSurv analysis on these data. Use a tau
> ## objective, with a minimum performance of 0.6. Note that
> ## messinaSurv analyses are very computationally-intensive,
> ## so in actual use multicore use with doMC and parallel = TRUE
> ## is strongly recommended.
> fit = messinaSurv(kirc.exprs, kirc.surv, obj_func = "tau", obj_min = 0.6)
Performance bootstrapping...
| | 0% |================= |33.33333% ~9 s remaining |================================== |66.66667% ~5 s remaining |====================================================|100% ~0 s remaining |====================================================|100% Completed after 16 s
Final training...
| | 0% |================= |33.33333% ~0 s remaining |================================== |66.66667% ~0 s remaining |====================================================|100% ~0 s remaining |====================================================|100% Completed after 0 s
>
> fit
An object of class MessinaSurvResult
Problem type:survival
Parameters:
An object of class MessinaParameters
3 features, 422 samples.
Objective type: survival (tau). Minimum objective value: 0.6
Minimum group fraction: 0.1
Training fraction: 0.8
Number of bootstraps: 50
Random seed:
Summary of results:
An object of class MessinaFits
1 / 3 features passed performance requirements (33.33%)
Top features:
Passed Requirements Classifier Type Threshold Value Direction
SAA1|6288 TRUE Threshold 5.207119 1
CCDC74A|90557 FALSE <NA> 7.617463 1
C1orf168|199920 FALSE <NA> NA NA
Margin
SAA1|6288 7.97642745
CCDC74A|90557 0.04505006
C1orf168|199920 0.00000000
> plot(fit)
>
>
>
>
>
> dev.off()
null device
1
>
|