|
Last data update: 2014.03.03
|
R: Plot the distribution of cluster sizes
| plotSizeDistribution | R Documentation |
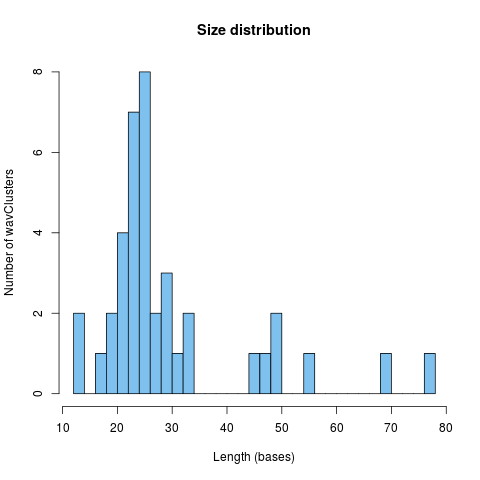
Plot the distribution of cluster sizes
Description
Produce an histogram of cluster sizes
Usage
plotSizeDistribution( clusters, showCov = FALSE, ... )
Arguments
clusters |
GRanges object containing individual clusters as identified
by the getClusters function
|
showCov |
logical, if TRUE a scatter plot of average cluster coverage
vs. cluster size is shown along with a loess fit. Default is FALSE.
|
... |
Additional parameters to be passed to the hist function
|
Value
Called for its effect, returns a histogram.
Author(s)
Federico Comoglio
See Also
getClusters
Examples
require(BSgenome.Hsapiens.UCSC.hg19)
data( model, package = "wavClusteR" )
filename <- system.file( "extdata", "example.bam", package = "wavClusteR" )
example <- readSortedBam( filename = filename )
countTable <- getAllSub( example, minCov = 10, cores = 1 )
highConfSub <- getHighConfSub( countTable, supportStart = 0.2, supportEnd = 0.7, substitution = "TC" )
coverage <- coverage( example )
clusters <- getClusters( highConfSub = highConfSub,
coverage = coverage,
sortedBam = example,
method = 'mrn',
cores = 1,
threshold = 2 )
fclusters <- filterClusters( clusters = clusters,
highConfSub = highConfSub,
coverage = coverage,
model = model,
genome = Hsapiens,
refBase = 'T',
minWidth = 12 )
plotSizeDistribution(fclusters, breaks = 30, col = 'skyblue2')
Results
R version 3.3.1 (2016-06-21) -- "Bug in Your Hair"
Copyright (C) 2016 The R Foundation for Statistical Computing
Platform: x86_64-pc-linux-gnu (64-bit)
R is free software and comes with ABSOLUTELY NO WARRANTY.
You are welcome to redistribute it under certain conditions.
Type 'license()' or 'licence()' for distribution details.
R is a collaborative project with many contributors.
Type 'contributors()' for more information and
'citation()' on how to cite R or R packages in publications.
Type 'demo()' for some demos, 'help()' for on-line help, or
'help.start()' for an HTML browser interface to help.
Type 'q()' to quit R.
> library(wavClusteR)
Loading required package: GenomicRanges
Loading required package: BiocGenerics
Loading required package: parallel
Attaching package: 'BiocGenerics'
The following objects are masked from 'package:parallel':
clusterApply, clusterApplyLB, clusterCall, clusterEvalQ,
clusterExport, clusterMap, parApply, parCapply, parLapply,
parLapplyLB, parRapply, parSapply, parSapplyLB
The following objects are masked from 'package:stats':
IQR, mad, xtabs
The following objects are masked from 'package:base':
Filter, Find, Map, Position, Reduce, anyDuplicated, append,
as.data.frame, cbind, colnames, do.call, duplicated, eval, evalq,
get, grep, grepl, intersect, is.unsorted, lapply, lengths, mapply,
match, mget, order, paste, pmax, pmax.int, pmin, pmin.int, rank,
rbind, rownames, sapply, setdiff, sort, table, tapply, union,
unique, unsplit
Loading required package: S4Vectors
Loading required package: stats4
Attaching package: 'S4Vectors'
The following objects are masked from 'package:base':
colMeans, colSums, expand.grid, rowMeans, rowSums
Loading required package: IRanges
Loading required package: GenomeInfoDb
Loading required package: Rsamtools
Loading required package: Biostrings
Loading required package: XVector
> png(filename="/home/ddbj/snapshot/RGM3/R_BC/result/wavClusteR/plotSizeDistribution.Rd_%03d_medium.png", width=480, height=480)
> ### Name: plotSizeDistribution
> ### Title: Plot the distribution of cluster sizes
> ### Aliases: plotSizeDistribution
> ### Keywords: graphics postprocessing
>
> ### ** Examples
>
>
> require(BSgenome.Hsapiens.UCSC.hg19)
Loading required package: BSgenome.Hsapiens.UCSC.hg19
Loading required package: BSgenome
Loading required package: rtracklayer
>
> data( model, package = "wavClusteR" )
>
> filename <- system.file( "extdata", "example.bam", package = "wavClusteR" )
> example <- readSortedBam( filename = filename )
> countTable <- getAllSub( example, minCov = 10, cores = 1 )
Loading required package: doMC
Loading required package: foreach
Loading required package: iterators
Considering substitutions, n = 497, processing in 1 chunks
chunk #: 1
considering the + strand
Computing local coverage at substitutions...
considering the - strand
Computing local coverage at substitutions...
> highConfSub <- getHighConfSub( countTable, supportStart = 0.2, supportEnd = 0.7, substitution = "TC" )
> coverage <- coverage( example )
> clusters <- getClusters( highConfSub = highConfSub,
+ coverage = coverage,
+ sortedBam = example,
+ method = 'mrn',
+ cores = 1,
+ threshold = 2 )
Computing start/end read positions
Number of chromosomes exhibiting high confidence transitions: 1
...Processing = chrX
>
> fclusters <- filterClusters( clusters = clusters,
+ highConfSub = highConfSub,
+ coverage = coverage,
+ model = model,
+ genome = Hsapiens,
+ refBase = 'T',
+ minWidth = 12 )
Computing log odds...
Refining cluster sizes...
Combining clusters...
Quantifying transitions within clusters...
Computing statistics...
| |== | 3% | |==== | 5% | |===== | 8% | |======= | 10% | |========= | 13% | |=========== | 15% | |============= | 18% | |============== | 21% | |================ | 23% | |================== | 26% | |==================== | 28% | |====================== | 31% | |======================= | 33% | |========================= | 36% | |=========================== | 38% | |============================= | 41% | |=============================== | 44% | |================================ | 46% | |================================== | 49% | |==================================== | 51% | |====================================== | 54% | |======================================= | 56% | |========================================= | 59% | |=========================================== | 62% | |============================================= | 64% | |=============================================== | 67% | |================================================ | 69% | |================================================== | 72% | |==================================================== | 74% | |====================================================== | 77% | |======================================================== | 79% | |========================================================= | 82% | |=========================================================== | 85% | |============================================================= | 87% | |=============================================================== | 90% | |================================================================= | 92% | |================================================================== | 95% | |==================================================================== | 97% | |======================================================================| 100%
Consolidating results...
> plotSizeDistribution(fclusters, breaks = 30, col = 'skyblue2')
>
>
>
>
>
>
> dev.off()
null device
1
>
|
|
 providing
providing  .
.