Supported by Dr. Osamu Ogasawara and  providing providing  . . |
|
Last data update: 2014.03.03 |
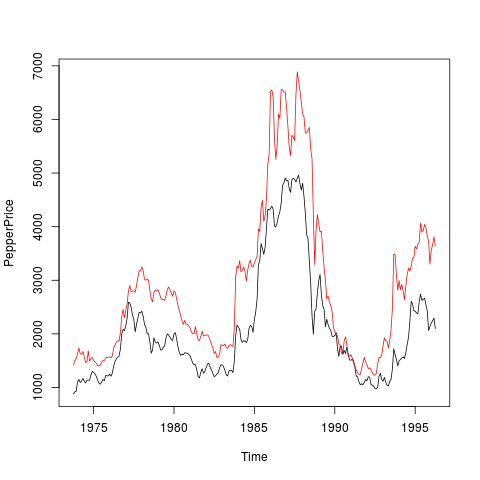
Black and White Pepper PricesDescriptionTime series of average monthly European spot prices for black and white pepper (fair average quality) in US dollars per ton. Usagedata("PepperPrice")
FormatA monthly multiple time series from 1973(10) to 1996(4) with 2 variables.
SourceOnline complements to Franses (1998). http://www.few.eur.nl/few/people/franses/research/book2.htm ReferencesFranses, P.H. (1998). Time Series Models for Business and Economic Forecasting. Cambridge, UK: Cambridge University Press. Examples
## data
data("PepperPrice")
plot(PepperPrice, plot.type = "single", col = 1:2)
## package
library("tseries")
library("urca")
## unit root tests
adf.test(log(PepperPrice[, "white"]))
adf.test(diff(log(PepperPrice[, "white"])))
pp.test(log(PepperPrice[, "white"]), type = "Z(t_alpha)")
pepper_ers <- ur.ers(log(PepperPrice[, "white"]),
type = "DF-GLS", model = "const", lag.max = 4)
summary(pepper_ers)
## stationarity tests
kpss.test(log(PepperPrice[, "white"]))
## cointegration
po.test(log(PepperPrice))
pepper_jo <- ca.jo(log(PepperPrice), ecdet = "const", type = "trace")
summary(pepper_jo)
pepper_jo2 <- ca.jo(log(PepperPrice), ecdet = "const", type = "eigen")
summary(pepper_jo2)
Results
R version 3.3.1 (2016-06-21) -- "Bug in Your Hair"
Copyright (C) 2016 The R Foundation for Statistical Computing
Platform: x86_64-pc-linux-gnu (64-bit)
R is free software and comes with ABSOLUTELY NO WARRANTY.
You are welcome to redistribute it under certain conditions.
Type 'license()' or 'licence()' for distribution details.
R is a collaborative project with many contributors.
Type 'contributors()' for more information and
'citation()' on how to cite R or R packages in publications.
Type 'demo()' for some demos, 'help()' for on-line help, or
'help.start()' for an HTML browser interface to help.
Type 'q()' to quit R.
> library(AER)
Loading required package: car
Loading required package: lmtest
Loading required package: zoo
Attaching package: 'zoo'
The following objects are masked from 'package:base':
as.Date, as.Date.numeric
Loading required package: sandwich
Loading required package: survival
> png(filename="/home/ddbj/snapshot/RGM3/R_CC/result/AER/PepperPrice.Rd_%03d_medium.png", width=480, height=480)
> ### Name: PepperPrice
> ### Title: Black and White Pepper Prices
> ### Aliases: PepperPrice
> ### Keywords: datasets
>
> ### ** Examples
>
> ## data
> data("PepperPrice")
> plot(PepperPrice, plot.type = "single", col = 1:2)
>
> ## package
> library("tseries")
> library("urca")
>
> ## unit root tests
> adf.test(log(PepperPrice[, "white"]))
Augmented Dickey-Fuller Test
data: log(PepperPrice[, "white"])
Dickey-Fuller = -1.744, Lag order = 6, p-value = 0.6838
alternative hypothesis: stationary
> adf.test(diff(log(PepperPrice[, "white"])))
Augmented Dickey-Fuller Test
data: diff(log(PepperPrice[, "white"]))
Dickey-Fuller = -5.336, Lag order = 6, p-value = 0.01
alternative hypothesis: stationary
Warning message:
In adf.test(diff(log(PepperPrice[, "white"]))) :
p-value smaller than printed p-value
> pp.test(log(PepperPrice[, "white"]), type = "Z(t_alpha)")
Phillips-Perron Unit Root Test
data: log(PepperPrice[, "white"])
Dickey-Fuller Z(t_alpha) = -1.6439, Truncation lag parameter = 5,
p-value = 0.726
alternative hypothesis: stationary
> pepper_ers <- ur.ers(log(PepperPrice[, "white"]),
+ type = "DF-GLS", model = "const", lag.max = 4)
> summary(pepper_ers)
###############################################
# Elliot, Rothenberg and Stock Unit Root Test #
###############################################
Test of type DF-GLS
detrending of series with intercept
Call:
lm(formula = dfgls.form, data = data.dfgls)
Residuals:
Min 1Q Median 3Q Max
-0.21135 -0.03069 -0.00108 0.03030 0.31018
Coefficients:
Estimate Std. Error t value Pr(>|t|)
yd.lag -0.004022 0.006015 -0.669 0.504
yd.diff.lag1 0.336267 0.061986 5.425 1.32e-07 ***
yd.diff.lag2 -0.105024 0.065414 -1.606 0.110
yd.diff.lag3 0.001263 0.065366 0.019 0.985
yd.diff.lag4 0.011251 0.062085 0.181 0.856
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
Residual standard error: 0.06481 on 261 degrees of freedom
Multiple R-squared: 0.1028, Adjusted R-squared: 0.0856
F-statistic: 5.98 on 5 and 261 DF, p-value: 2.947e-05
Value of test-statistic is: -0.6686
Critical values of DF-GLS are:
1pct 5pct 10pct
critical values -2.57 -1.94 -1.62
>
> ## stationarity tests
> kpss.test(log(PepperPrice[, "white"]))
KPSS Test for Level Stationarity
data: log(PepperPrice[, "white"])
KPSS Level = 0.91295, Truncation lag parameter = 3, p-value = 0.01
Warning message:
In kpss.test(log(PepperPrice[, "white"])) :
p-value smaller than printed p-value
>
> ## cointegration
> po.test(log(PepperPrice))
Phillips-Ouliaris Cointegration Test
data: log(PepperPrice)
Phillips-Ouliaris demeaned = -24.099, Truncation lag parameter = 2,
p-value = 0.02404
> pepper_jo <- ca.jo(log(PepperPrice), ecdet = "const", type = "trace")
> summary(pepper_jo)
######################
# Johansen-Procedure #
######################
Test type: trace statistic , without linear trend and constant in cointegration
Eigenvalues (lambda):
[1] 4.931953e-02 1.350807e-02 2.081668e-17
Values of teststatistic and critical values of test:
test 10pct 5pct 1pct
r <= 1 | 3.66 7.52 9.24 12.97
r = 0 | 17.26 17.85 19.96 24.60
Eigenvectors, normalised to first column:
(These are the cointegration relations)
black.l2 white.l2 constant
black.l2 1.0000000 1.00000 1.000000
white.l2 -0.8892307 -5.09942 2.280911
constant -0.5569943 33.02742 -20.032441
Weights W:
(This is the loading matrix)
black.l2 white.l2 constant
black.d -0.07472300 0.002453210 -4.958157e-18
white.d 0.02015611 0.003537005 8.850353e-18
> pepper_jo2 <- ca.jo(log(PepperPrice), ecdet = "const", type = "eigen")
> summary(pepper_jo2)
######################
# Johansen-Procedure #
######################
Test type: maximal eigenvalue statistic (lambda max) , without linear trend and constant in cointegration
Eigenvalues (lambda):
[1] 4.931953e-02 1.350807e-02 2.081668e-17
Values of teststatistic and critical values of test:
test 10pct 5pct 1pct
r <= 1 | 3.66 7.52 9.24 12.97
r = 0 | 13.61 13.75 15.67 20.20
Eigenvectors, normalised to first column:
(These are the cointegration relations)
black.l2 white.l2 constant
black.l2 1.0000000 1.00000 1.000000
white.l2 -0.8892307 -5.09942 2.280911
constant -0.5569943 33.02742 -20.032441
Weights W:
(This is the loading matrix)
black.l2 white.l2 constant
black.d -0.07472300 0.002453210 -4.958157e-18
white.d 0.02015611 0.003537005 8.850353e-18
>
>
>
>
>
> dev.off()
null device
1
>
|