Supported by Dr. Osamu Ogasawara and  providing providing  . . |
|
Last data update: 2014.03.03 |
Soil Electrical ConductivityDescriptionElectrical conductivity of soil paste extracts from the Lower Arkansas River Valley, at sites upstream and downstream of the John Martin Reservoir. Usagedata(ArkansasRiver) FormatThe format is: List of 2 $ upstream : num [1:823] 2.37 3.53 3.06 3.35 3.07 ... $ downstream: num [1:435] 8.75 6.59 5.09 6.03 5.64 ... DetailsElectrical conductivity is a measure of soil water salinity. SourceThis data set was supplied by Eric Morway (emorway@usgs.gov). ReferencesEric D. Morway and Timothy K. Gates (2011) Regional assessment of soil water salinity across an extensively irrigated river valley. Journal of Irrigation and Drainage Engineering, doi:10.1061/(ASCE)IR.1943-4774.0000411 Examples
data(ArkansasRiver)
lapply(ArkansasRiver, summary)
upstream <- ArkansasRiver[[1]]
downstream <- ArkansasRiver[[2]]
## Fit normal inverse Gaussian
## Hyperbolic can also be fitted but fit is not as good
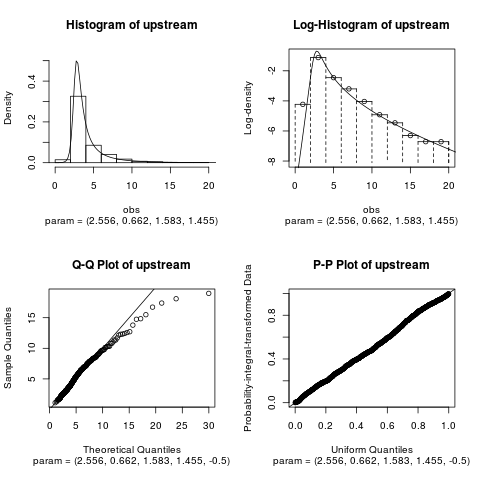
fitUpstream <- nigFit(upstream)
summary(fitUpstream)
par(mfrow = c(2,2))
plot(fitUpstream)
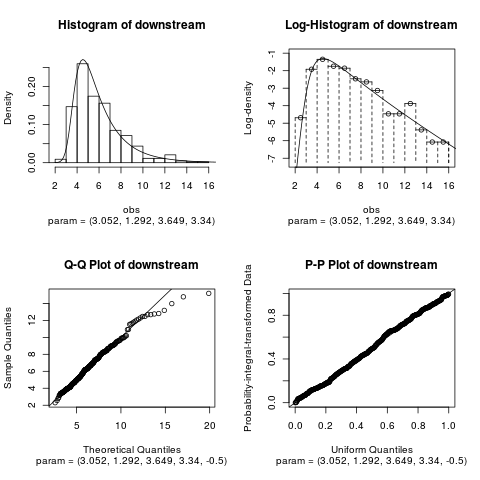
fitDownstream <- nigFit(downstream)
summary(fitDownstream)
plot(fitDownstream)
par(mfrow = c(1,1))
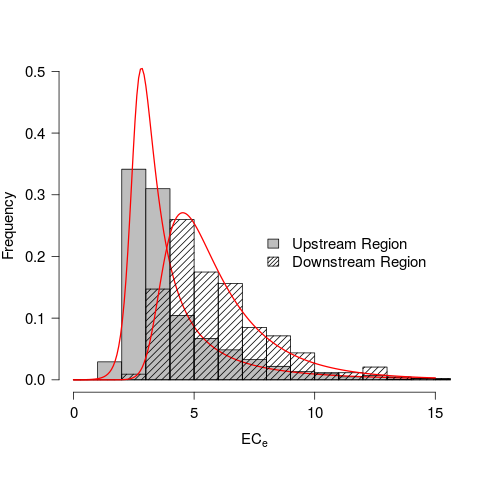
## Combined plot to compare
## Reproduces Figure 3 from Morway and Gates (2011)
hist(upstream, col = "grey", xlab = "", ylab = "", cex.axis = 1.25,
main = "", breaks = seq(0,20, by = 1), xlim = c(0,15), las = 1,
ylim = c(0,0.5), freq = FALSE)
param <- coef(fitUpstream)
nigDens <- function(x) dnig(x, param = param)
curve(nigDens, 0, 15, n = 201, add = TRUE,
ylab = NULL, col = "red", lty = 1, lwd = 1.7)
hist(downstream, add = TRUE, col = "black", angle = 45, density = 15,
breaks = seq(0,20, by = 1), freq = FALSE)
param <- coef(fitDownstream)
nigDens <- function(x) dnig(x, param = param)
curve(nigDens, 0, 15, n = 201, add = TRUE,
ylab = NULL, col = "red", lty = 1, lwd = 1.7)
mtext(expression(EC[e]), side = 1, line = 3, cex = 1.25)
mtext("Frequency", side = 2, line = 3, cex = 1.25)
legend(x = 7.5, y = 0.250, c("Upstream Region","Downstream Region"),
col = c("black","black"), density = c(NA,25),
fill = c("grey","black"), angle = c(NA,45),
cex = 1.25, bty = "n", xpd = TRUE)
Results
R version 3.3.1 (2016-06-21) -- "Bug in Your Hair"
Copyright (C) 2016 The R Foundation for Statistical Computing
Platform: x86_64-pc-linux-gnu (64-bit)
R is free software and comes with ABSOLUTELY NO WARRANTY.
You are welcome to redistribute it under certain conditions.
Type 'license()' or 'licence()' for distribution details.
R is a collaborative project with many contributors.
Type 'contributors()' for more information and
'citation()' on how to cite R or R packages in publications.
Type 'demo()' for some demos, 'help()' for on-line help, or
'help.start()' for an HTML browser interface to help.
Type 'q()' to quit R.
> library(GeneralizedHyperbolic)
Loading required package: DistributionUtils
Loading required package: RUnit
> png(filename="/home/ddbj/snapshot/RGM3/R_CC/result/GeneralizedHyperbolic/ArkansasRiver.Rd_%03d_medium.png", width=480, height=480)
> ### Name: ArkansasRiver
> ### Title: Soil Electrical Conductivity
> ### Aliases: ArkansasRiver
> ### Keywords: datasets
>
> ### ** Examples
>
> data(ArkansasRiver)
> lapply(ArkansasRiver, summary)
$upstream
Min. 1st Qu. Median Mean 3rd Qu. Max.
1.107 2.767 3.255 4.104 4.602 18.980
$downstream
Min. 1st Qu. Median Mean 3rd Qu. Max.
2.296 4.360 5.361 5.986 7.009 15.180
> upstream <- ArkansasRiver[[1]]
> downstream <- ArkansasRiver[[2]]
> ## Fit normal inverse Gaussian
> ## Hyperbolic can also be fitted but fit is not as good
> fitUpstream <- nigFit(upstream)
> summary(fitUpstream)
Data: upstream
Parameter estimates:
mu delta alpha beta
2.5557 0.6616 1.5825 1.4551
Likelihood: -1441.33
Method: Nelder-Mead
Convergence code: 0
Iterations: 223
> par(mfrow = c(2,2))
> plot(fitUpstream)
> fitDownstream <- nigFit(downstream)
> summary(fitDownstream)
Data: downstream
Parameter estimates:
mu delta alpha beta
3.052 1.292 3.649 3.340
Likelihood: -880.0715
Method: Nelder-Mead
Convergence code: 0
Iterations: 293
> plot(fitDownstream)
> par(mfrow = c(1,1))
> ## Combined plot to compare
> ## Reproduces Figure 3 from Morway and Gates (2011)
> hist(upstream, col = "grey", xlab = "", ylab = "", cex.axis = 1.25,
+ main = "", breaks = seq(0,20, by = 1), xlim = c(0,15), las = 1,
+ ylim = c(0,0.5), freq = FALSE)
> param <- coef(fitUpstream)
> nigDens <- function(x) dnig(x, param = param)
> curve(nigDens, 0, 15, n = 201, add = TRUE,
+ ylab = NULL, col = "red", lty = 1, lwd = 1.7)
>
> hist(downstream, add = TRUE, col = "black", angle = 45, density = 15,
+ breaks = seq(0,20, by = 1), freq = FALSE)
> param <- coef(fitDownstream)
> nigDens <- function(x) dnig(x, param = param)
> curve(nigDens, 0, 15, n = 201, add = TRUE,
+ ylab = NULL, col = "red", lty = 1, lwd = 1.7)
>
> mtext(expression(EC[e]), side = 1, line = 3, cex = 1.25)
> mtext("Frequency", side = 2, line = 3, cex = 1.25)
> legend(x = 7.5, y = 0.250, c("Upstream Region","Downstream Region"),
+ col = c("black","black"), density = c(NA,25),
+ fill = c("grey","black"), angle = c(NA,45),
+ cex = 1.25, bty = "n", xpd = TRUE)
>
>
>
>
>
> dev.off()
null device
1
>
|